Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- Conclusion
- References
- Copyright
Analysis of Blood Samples to Identify Leukemia Cells using a Support Vector Machine and K-Nearest Neighbor Algorithms
Authors: Savithri ., Santhosh , Rajkumar , Sreerambabu
DOI Link: https://doi.org/10.22214/ijraset.2023.54994
Certificate: View Certificate
Abstract
The study proposes an innovative approach using MATLAB to automate the counting of leukemia cells in blood samples, employing Support Vector Machine (SVM) and Nearest Neighbor algorithms. The method involves preprocessing blood sample images to enhance contrast and apply image filters, followed by segmentation techniques for isolating individual cells. SVM and nearest neighbor algorithms are trained using extracted features such as cell size, shape, and texture. Accurate detection and counting of leukemia cells play a crucial role in leukemia diagnosis and management. Leukemia is a group of cancers characterized by abnormal white blood cell proliferation in the bone marrow, leading to symptoms like bleeding, bruising, fatigue, and increased infection risk due to insufficient normal blood cells. Diagnosis typically involves blood tests or bone marrow biopsy. In clinical bioinformatics, SVM algorithms have enabled the development of robust experimental cancer diagnostic models, utilizing gene expression data with a small number of samples and numerous variables. Efficient implementations of SVM algorithms further facilitate practical application. Support Vector Machines excel in mapping data to higher-dimensional spaces through kernel functions, allowing the identification of maximum-margin hyperplanes for separating training instances.
Introduction
I. INTRODUCTION
The classification of medical images by various algorithms and classifiers has seen significant growth, leading to improved accuracy and aiding in the diagnosis for healthcare students. Effective medical images play a crucial role in supporting their studies. Data mining involves extracting meaningful correlations, patterns, and trends from large databases. Initially, algorithms were developed to identify natural grouping tendencies in data, relying on mathematical statistics and heuristic graphical techniques. These systems have enhanced the accuracy of the classification process.
Biomedical image processing encompasses various techniques, algorithms, and classifiers for medical image classification, which have revolutionized visualization and interpretation in biology and medicine.
The field requires close collaboration between physicians and engineers to design, implement, and validate integrated medical systems.
The primary objective of image analysis is to gather information, detect diseases, diagnose conditions, provide control and therapy, and monitor and evaluate progress. Currently, blood disorders are identified through visual inspection of microscopic blood cell images, which can aid in the classification of related diseases. Among them, leukemia is a particularly dangerous form of blood cancer, and late detection can have fatal consequences.
The proposed approach, implementing SVM and nearest neighbor algorithms in MATLAB, shows promise for accurate leukemia diagnosis and management.
Leveraging advanced image processing techniques and machine learning algorithms within MATLAB significantly enhances the accuracy and efficiency of leukemia cell counting, providing a reliable tool for medical professionals. This paper introduces a novel method for effectively segmenting various types of leukemia cells, such as monocytes, lymphocytes, eosinophils, basophils, and neutrophils, from microscopic blood images. The segmentation technique utilizes the HSV saturation component and incorporates blob analysis to achieve precise cell isolation. Moreover, integrating SVM and KNN algorithms ensures improved counting accuracy, yielding more reliable results.
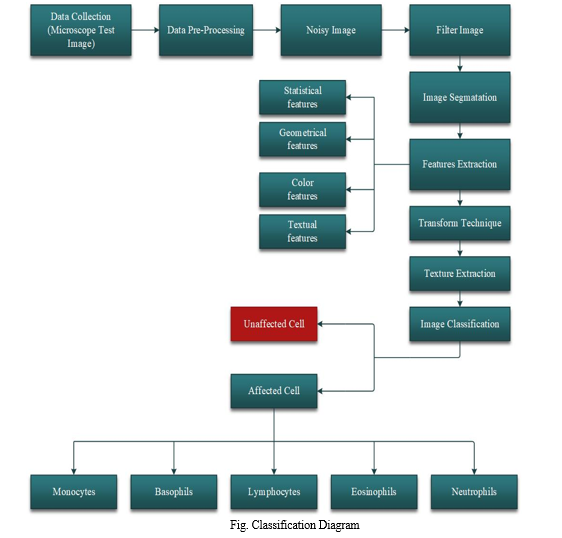
II. PROPOSED METHODOLOGY
A proposed MATLAB system incorporates image processing techniques and machine learning algorithms to automate the detection and classification of leukemia cells in blood smears. The system aims to analyze blood samples and count leukemia cells using a support vector machine and k-nearest neighbor algorithms. Through the utilization of classification techniques, such as nearest neighbor and SVM, diseases related to blood can be detected by identifying and counting blood cells within the blood smear. The system is designed to enhance efficiency, accuracy, and reliability in leukemia cell counting, providing clinicians with timely and precise diagnostic information.
The system's three key components include image pre-processing, feature extraction, and classification, with digital images of blood smears serving as input after being captured through a microscope or digital camera.
These images undergo pre-processing to enhance quality, eliminate noise, and segment individual cells. The Support Vector Machine method leverages the mapping of data to a higher dimensional space using a kernel function, enabling the identification of a maximum-margin hyperplane that separates training instances. On the other hand, the K-nearest neighbors (KNN) algorithm treats samples as points in a multidimensional space. It classifies unseen models based on a vote from the K closest training instances, determined by a distance metric like Euclidean distance.
A. Support Vector Machines (SVM)
The support vector machine (SVM) is a classification algorithm widely used in various domains such as handwriting recognition, object recognition, speaker identification, face detection, and text categorization. In computational biology, SVMs have shown exceptional performance, enabling the development of powerful cancer diagnostic models using gene expression data, even with limited samples and a large number of variables. Furthermore, efficient and high-quality implementations of SVM algorithms make their practical application feasible. Initially, SVMs were limited to binary classification tasks.
Preliminary experimental evidence suggests that certain multi-category SVMs (MCSVMs) exhibit strong performance in cancer diagnostic experiments based on gene expression data. In the following method descriptions, "k" represents the number of diagnostic categories or classes, while "n" represents the number of samples or patients in the training dataset.
B. K Nearest Neighbors (KNN)
The fundamental concept behind K Nearest Neighbors (KNN) is to consider all samples as points in an m-dimensional space, where "m" corresponds to the number of variables. When presented with an unseen sample "x," the algorithm classifies it by tallying the votes from the K nearest training instances, determined by a distance metric, often using Euclidean distance.
The research methodology employed in this study includes the following steps:
- Image Acquisition: Blood cell images will be obtained from a nearby hospital using either effective enlargement techniques or a digital microscope.
2. Image Pre-processing: The pre-processing tasks aim to enhance image quality and remove overlapping blood cells in the image's border area. This involves several subtasks, including:
- Extraction of the green plane: The green plane is extracted from the blood cell image since it contains the most relevant information.
- Histogram equalization: Intensity values of the image are adjusted through histogram equalization, aligning the histogram with a predefined distribution.
- Contrast and brightness adjustment: The image's contrast and brightness are adjusted based on the histogram and the desired display range.
3. Image Segmentation: Various segmentation methods, such as threshold-based, edge-based, region-based, or clustering methods (e.g., fuzzy-C mean clustering, K-means clustering), have been proposed. The chosen method in this research is described as follows:
- Conversion of the color blood slide image to grayscale.
- Contrast enhancement using histogram equalization.
- Linear contrast stretching to adjust image intensity levels.
- Creation of multiple images to highlight different components.
- Application of a minimum filter to reduce noise and preserve edges.
- Global thresholding using Otsu's method.
- Conversion to a binary image and removal of small pixel groups through morphological opening.
- Connection of neighboring pixels to form objects.
- Removal of objects smaller than 50% of the average RBC area using a size test.
4. Feature Extraction: The classification task involves assigning an unknown test vector to a known class. In this step, a reinforcement learning algorithm is proposed to classify different types of leukemia, including ALL, AML, CLL, and CML. The area of the image region was calculated by counting the total number of non-zero pixels present. The perimeter was measured by calculating the distance between consecutive boundary pixels. Circularity, a dimensionless parameter, represents the surface irregularities and is computed using the formula:
Circularity = 4 * Pi * Area / Perimeter^2.
5. Image classification: In the image classification step, the extracted features are utilized to classify lymphocyte cells as either blast or normal cells. Classification involves assigning an unknown test vector to a known class label. The k-nearest neighbor (KNN) decision rule is commonly used for classification due to its scalability and effectiveness. Selecting the optimal value of k is often a laborious task as it depends on the specific dataset. The k-nearest neighbors (KNN) algorithm is a non-parametric classification method that is simple yet highly effective in many scenarios. In this study, KNN is employed to classify blast cells from normal white blood cells.
III. CLASSIFICATION AND RECOGNITION
The primary objective of this study is to employ machine learning techniques and an image dataset for the purpose of detecting different types of WBCs. The forthcoming sections will offer a comprehensive overview of WBC, furnish details regarding the dataset, and delve into the intricacies of feature engineering and model selection.
A. White Blood Cells
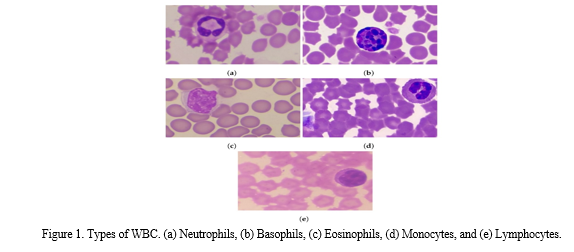
White blood cells (WBCs) lack pigmentation, resulting in their colorless appearance within the bloodstream. In a cubic mm of blood, there are typically 7000 to 8000 WBCs, which are considerably larger than red blood cells. WBCs can be classified into five distinct types based on their nucleus shape and cytoplasmic granule density. Notably, WBCs stand out from other blood cells due to their possession of a single large bilobed nucleus. These specialized cells originate in the bone marrow and subsequently migrate to the blood and lymphatic system. Here is a concise overview of each WBC type:
- Neutrophils represent around 62% of the total WBC count. They exhibit the remarkable ability to engulf and neutralize foreign particles, including viruses and bacteria, thereby counteracting their harmful effects. Neutrophils are approximately twice the size of red blood cells and feature a nucleus containing 2 to 5 lobes.
- Basophils account for less than 1% of WBCs. They are roughly twice the size of red blood cells and possess a bilobed nucleus. Basophils release heparin, a protein that prevents blood clotting, and secrete histamine, which induces inflammation. Additionally, they release antibodies and antibiotics to shield the body against the impact of foreign substances.
- Monocytes make up approximately 3% of the total WBC count. They are roughly two to three times larger than red blood cells and have a nucleus that ranges from nearly round to lobed in shape. Monocytes give rise to macrophages, which participate in phagocytosis to eliminate larger particles. Macrophages have a lifespan of approximately 8 to 10 hours in the blood before migrating to lymphoid tissue, where they transform into macrophages.
- Eosinophils constitute about 2% of WBCs. They are approximately twice the size of red blood cells and possess a bilobed nucleus. Eosinophils play a role in deactivating substances that promote inflammation and engage in combatting parasites and worms.
- Lymphocytes make up approximately 32% of the total WBC count. Their size closely resembles that of red blood cells. Lymphocytes are responsible for antibody production. The lifespan of lymphocytes in the blood can vary, lasting from months to years depending on the level of their activity.
IV. FLOW OF IMPLEMENTED METHODOLOGY
In this study, we obtained a dataset of WBC images from the IEEE data port, which was used for feature extraction. Two types of features, namely texture features and RGB features, were extracted. The skimage library was utilized for extracting the texture features. Following feature extraction, a feature selection technique using Chi2 was employed to select the most prominent and essential features. From both the RGB and texture features, 3000 important features were selected. These selected features from both categories were combined to create a hybrid feature set. To address the issue of an imbalanced dataset, we applied data oversampling using SMOTE. This approach helps to mitigate overfitting of the model towards the majority class. For WBC-type classification, various machine learning and deep learning models were employed. The dataset was split into training and testing sets with a ratio of 0.8 for training and 0.2 for testing. The performance of all models was evaluated based on accuracy, precision, recall, and F1 score.
V. EXPERIMENTAL RESULT
The proposed technique was applied to analyze a set of 121 peripheral blood smear images obtained from the previously mentioned public dataset. An evaluation was performed using a microscopic blood image with dimensions of 2592 × 1944 pixels. In addition to the analysis, the number of white blood cells (WBCs) present in each image was also determined.
The process involved separating WBCs from other blood components by identifying white spots over the nuclei. Subsequently, in the final segmented image, only the WBCs with darker nuclei were retained, while noisy components were eliminated using a minimum filter. Following this segmentation, lymphocyte cells were identified within the WBC population using blast segmentation, and the morphological features of lymphocytes were calculated. Based on these features, a decision was made to determine whether the slide image indicated the presence of leukemia or not.
VI. ACKNOWLEDGEMENT
I acknowledge our mentor Mr. N. Santhosh who provided insight and expertise that greatly helped the research, for suggestions that greatly improved this manuscript. Special thanks, to our supervisor Mr. M. Mohammed Riyaz for the support in this research work.
Conclusion
The project focuses on the detection of leukemia cell types using microscopic blood sample images. The system utilizes various features, including texture, geometry, colors, and statistical analysis, as inputs for the classifier. The system aims to be efficient, and reliable, have a shorter processing time, lower error rate, high accuracy, cost-effectiveness, and robustness towards individual variations, sample collection protocols, and time variations. The true strength of Support Vector Machines lies in mapping the data to a higher dimensional space through a kernel function, enabling the identification of a maximum-margin hyperplane that separates training instances. By employing the proposed features, leukemia detection achieved an overall accuracy of 93% using the K-nearest neighbors (KNN) classifier. Additionally, the system should be capable of handling excessive staining and cell overlap challenges. Enhancements refer to significant improvements made to the product, either as part of a new version or to enhance existing capabilities. The project holds immense potential for future development. The system should also exhibit robustness in the presence of excessive staining and overlapping cells. The obtained results provide encouragement for future endeavors, including the classification of lymphoblasts into different subtypes. Exploring alternative techniques for stain-independent blood smear image segmentation and leukemia-type classification is also recommended. The project can be easily updated in the future to meet specific requirements, as it offers flexibility for expansion. With the implementation of the proposed database Space Manager software, the client can effectively manage and execute their work with enhanced accuracy and reduced errors.
References
[1] T. Rosyadi, A. Arif, Nopriadi, B. Achmad and Faridah, \"Classification of Leukocyte Images Using K-Means Clustering Based on Geometry Features,\" in 6th International Annual Engineering Seminar (InAES), Yogyakarta, Indonesia, 2016. [2] N. M. Salem, \"Segmentation of White Blood Cells from Microscopic Images using K-means clustering,\" in 2014 31st National Radio Science Conference (NRSC), 2014. [3] A. Gautam and H. Bhadauria, \"White Blood Nucleus Extraction Using K-Mean Clustering and Mathematical Morphing,\" in 5th International Conference- Confluence the Nect Generation Information Technology Summit (Confluence), 2014. [4] O. Ryabchykov, A. Ramoji, T. Bocklitz, M. Foerster, S. Hagel, C. Kroegel, M. Bauer, U. Neugebauer and J. Popp, \"Leukocyte subtypes classification by means of image processing,\" Proceedings of the Federal Conference on Computer Science and Information Systems, vol. 8, no. 2300-5963, pp. 309-316, 2016. [5] N. Sinha and A. G. Ramakrishnan, \"Automation of differential blood count,\" TENCON 2003. Conference on Convergent Technologies for Asia-Pacific Region, vol. 2, pp. 547-551, 2003. [6] D.-C. Huang and K.-D. Hung, \"Leukocyte Nucleus Segmentation and Recognition in Color Blood-Smear Images,\" in IEEE International Instrumentation and Measurement Technology, 2012. [7] Puttamadegowda and Prasannakumar, \"White Blood cel segmentation using Fuzzy C means and snake,\" in 2016 International Conference on Computation System and Information Technology for Sustainable Solutions (CSITSS), 2016. [8] R. Ahasan, A. U. Ratul and A. S. M. Bakibillah, \"White Blood Cells Nucleus Segmentation from Microscopic Images of strained peripheral blood film during Leukemia and Normal Condition,\" in 5th International Conference on Informatics, Electronics and Vision (ICIEV), 2016. [9] Z. K. K. Alreza and A. Karimian, \"Design a new algorithm to count white blood cells for classification Leukemic Blood Image using machine vision system,\" in International Conference on Computer and Knowledge Engineering (ICCKE 2016), 2016. [10] W. Yu, C. Yang, L. Zhang, H. Shen, Y. Xia and J. Sha, \"Automatic Classification of Leukocytes Using Deep Neural Network,\" in IEEE 12th International Conference on ASIC (ASICON), 2017.
Copyright
Copyright © 2023 Savithri ., Santhosh , Rajkumar , Sreerambabu . This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET54994
Publish Date : 2023-07-25
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

