Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- Conclusion
- References
- Copyright
Automated System for Medicinal Plant Classification Usage Recommendation
Authors: Hemalatha ., Sreerambabu , Rajkumar , Santhosh
DOI Link: https://doi.org/10.22214/ijraset.2023.55008
Certificate: View Certificate
Abstract
We propose the utilization of deep learning to create an automated system for classifying medicinal plants, enabling swift identification of beneficial plant species. Convolutional neural networks (CNNs) are renowned for their exceptional capabilities in extracting and classifying features. Our proposed model, named \"deep learning model,\" is developed by extracting features and employing a CNN classifier, resulting in high accuracy and minimal prediction time when applied to real-time images. The system consists of three modules: image pre-processing, image feature extraction, and image classification. During the initial pre-processing step, we conduct RGB conversion to extract the green band from input images. In the subsequent step, after pre-processing, we extract features such as shape, color, and texture from the pre-processed image. These features are then utilized to classify the image as either medicinal or a regular leaf. Plants have been utilized as a medicinal resource in Ayurveda since ancient times. The identification of the correct plant is a crucial step in the preparation of Ayurvedic medicine, traditionally carried out manually. However, with the need for mass production, it becomes essential to automate the process of plant identification. In this study, we propose the implementation of a technique that employs a CNN algorithm, along with an ensemble supervised machine learning algorithm that considers color, texture, and geometrical features, for the identification of medicinal plants. The Convolutional Neural Network (CNN) is utilized for classification and determining the plant class.
Introduction
I. INTRODUCTION
The primary objective of this project is to achieve accurate and fast detection and classification of medicinal plant leaf images. The project introduces two innovative approaches. Firstly, a deep learning model is constructed for the classification of medicinal herbs. Secondly, a custom dataset consisting of 500 images is created to identify 10 species of Indian (Ayurveda) herbs. The scope of a dedicated website focused on plant leaf medicine details and identification is to provide users with reliable and precise information about various types of plant leaves that possess medicinal properties, including their benefits and potential side effects.
Such a website could incorporate a tool that enables users to upload images of plant leaves and obtain information about the specific leaf type along with its medicinal properties. The platform would offer comprehensive details about the therapeutic benefits of different plant leaves, including the ailments they can treat and recommended dosages. Emphasizing the importance of user safety, the website would also provide information on potential side effects of using plant leaves for medicinal purposes, accompanied by recommendations on minimizing the risk of adverse reactions.
A medicinal plant refers to any plant that contains substances in one or more of its organs which can be utilized for therapeutic purposes or as precursors for valuable drugs. The term "medicinal plant" encompasses various plant types used in herbalism or heal medicine. Herbalism involves the use of plants for medicinal purposes and the study thereof. Plants such as Aloe, Tulsi, Neem, Turmeric, and Ginger possess the ability to treat numerous common ailments. These plants are commonly employed as home remedies across many regions in the country. It is widely recognized that Basil (Tulsi) is extensively used by consumers for medicinal preparations, black tea, religious ceremonies, and other daily activities.
II. PROPOSED METHODOLOGY
The identification of medicinal plants, a CNN classifier is utilized. Given that our proposed approach involves classifying multiple classes, we have expanded the binary CNN model to incorporate multinomial logistic regression. The RPN module serves the purpose of selecting numerous areas from the features map that potentially contain objects. It generates predictions for these regions, identifying those likely to contain objects and adjusting the frame size accordingly. Among the RPN predictions, the one with the highest foreground score is retained, while the remaining predictions are discarded.
To investigate the images, GLCM parameters were computed for each color component. The parameters, namely contrast, correlation, energy, and homogeneity, capture the texture's coarseness, linear dependency, textural uniformity, and pixel distribution, respectively CNN models operate under the assumption that the input images are converted into a single feature vector, which is then processed through a sequence of hidden layers. These layers consist of neurons arranged in three dimensions: width, height, and depth. Neurons in CNNs are connected to a specific region of the layer above them, rather than being connected to all neurons in a fully connected manner. Every layer in the CNN transforms the 3D volume input into 3D volume output activations.
The convolutional layer utilizes a filter (referred to as a kernel) to identify patterns in the input images (original image). Multiple filters can be used to extract different features. The filter is relatively small in size, enabling it to scan the entire image and perform specific arithmetic operations between the filter and the pixels to extract the desired features. The pooling layer is employed to reduce the size of the activation maps. While not essential, this step helps prevent overfitting. The underlying concept of pooling is straightforward, as it involves scaling down large arrays to a smaller size.
The fully-connected layer serves as the final layer, where each neuron is connected to all nodes in the previous layer. It is in this layer that the final classification process occurs. When designing the network model, an image is initially fed into a convolutional layer, and an activation fun ction, such as ReLU, is applied to the convolutional layer's output. The resulting output is then passed through additional convolutional layers, repeating this process several times until reaching the fully-connected layer. Each neuron in the convolutional layer acts as a filter, responsible for detecting various image features such as edges, curves, colors, textures, and density, based on the depth of the network.
III. RECOGINITION OF MEDICINAL PLANT
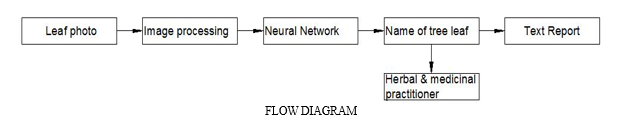
A. Image Acquisition
The initial step in the identification process involves obtaining an image of the plant. The image can be captured for various plant parts, such as the entire plant, leaf, flower, stem, or fruits. According to the authors, they categorize the images into three categories based on the acquisition method: 'scans', 'pseudo-scans', and 'photos'. In the scan and pseudo-scan categories, the leaf images are obtained either through scanning or photography, respectively. These images are captured indoors, in front of a plain background. The researchers predominantly utilize scan and pseudo-scan images due to their ease of examination. Typically, simple, fully grown, and unaltered leaves are selected for imaging, conducted in a well-lit laboratory setting. The utilization of scans and pseudo-scans simplifies the classification task since the images are captured against a plain background.
B. Leaf Image Preprocessing
Image pre-processing plays a crucial role in enhancing the quality of the image for subsequent processing steps. It is necessary because images typically contain noise, which can adversely affect classification accuracy. The purpose of pre-processing is to eliminate noise that may impede the identification process and address degraded data. To improve the leaf image, a series of operations are applied. These operations include converting the RGB image to grayscale, followed by further conversion from grayscale to binary. Additionally, techniques such as smoothing, filtering, and other relevant processes are employed. In this paper, the pre-processing mechanism encompasses noise reduction, resizing operations, and image enhancement, ensuring the improvement of the image quality for subsequent analysis.
Gaussian Filtering: In order to address image noise, this study utilizes Gaussian Filtering, also known as Gaussian Smoothing. Gaussian Filtering is a linear filter that effectively reduces noise and eliminates redundant information present in the image.
Resizing: Following the noise handling process, the images undergo a resizing operation. In this study, the images are resized to a dimension of [300 × 400].
Image Enhancement: Given that we are working with color images, image enhancement becomes a crucial step in our process. Furthermore, as the subsequent step involves color image segmentation, it is necessary to enhance the image's contrast and texture to achieve optimal results. Image enhancement aims to eliminate any redundant pixels in the color image prior to segmentation. In our study, we enhance the image's contrast through a technique called contrast stretching. This process improves the contrast within the image by expanding its dynamic range of intensity values. Additionally, contrast adjustment is applied, saturating the top one percent and bottom one percent of all pixel values. These steps collectively enhance the contrast and improve the overall quality of the image before proceeding with segmentation.
C. Leaf Vein Analysis
- Region Proposal Network: The Region Proposal Network (RPN) performs the task of selecting numerous areas from the features map that potentially contain objects. These selected regions are referred to as anchors, which serve as bounding boxes for the objects. In practical scenarios, there are a large number of anchors, often exceeding 200, with varying sizes and aspect ratios. The goal is for these anchors to cover the objects in the image as comprehensively as possible. The RPN prediction then identifies the anchors that are likely to contain the objects of interest and adjusts the frame size accordingly to encompass them. In cases where multiple anchors overlap excessively within a particular region, the RPN prediction retains the anchor with the highest foreground score and discards the remaining overlapping anchors.
- ROI Align: Utilizing bilinear interpolation, the ROL Align module can crop a specific region of interest (ROI) from the feature map. This cropped region is then passed on to the ROI classifier and bounding box regressor for further processing.
- Box Regression and Classification: In this stage, the ROI obtained from the ROL Align module is inputted. It consists of both the ROI classifier and bounding box regressor. The ROI classifier, which is a deeper network, refines the classification of the ROI. However, in this particular experiment, the ROI classifier is solely utilized for classifying two classes, namely foreground and background. On the other hand, the bounding box regressor functions similarly to the RPN, but with the additional capability of further refining the bounding box to better fit the object.
D. Feature Extraction
After the completion of pre-processing and segmentation to isolate the desired region of interest, the next crucial step is feature extraction. Feature extraction is recognized as one of the pivotal stages in image processing and pattern analysis. It can be viewed as a dimensionality reduction process. An image inherently contains a vast amount of information, not all of which can be efficiently processed. Additionally, handling such a large volume of data requires substantial computational power and memory resources. Consequently, feature extraction is performed to decrease the number of variables involved in the subsequent image processing stages. The selection of the appropriate set of features to accurately describe the image is of utmost importance.
- Texture Feature: Texture analysis holds significant importance in various domains, including medical imaging and image retrieval. Texture, within the context of image processing, encompasses properties such as smoothness, coarseness, and regularity. It characterizes the spatial distribution of grayscale levels within a neighborhood of pixels in a digital image. Texture feature extraction methods can be classified into four categories: statistical, structural, model-based, and transform-based. In our study, we focus on the statistical method, which characterizes texture by analyzing the statistical properties of the grayscale image.
- Colour Feature: During the segmentation phase, the input image was partitioned into three distinct color channels. Color features are then extracted individually from the resulting segmented images. In this study, the extracted color features can be considered as color-based texture features, as they are derived from first-order statistics calculated from the colored images themselves, rather than the conventional grayscale images.
E. CNN Leaf Classification
During the segmentation phase, the input image was partitioned into three distinct color channels. Color features are then extracted individually from the resulting segmented images. In this study, the extracted color features can be considered as color-based texture features, as they are derived from first-order statistics calculated from the colored images themselves, rather than the conventional grayscale images. During the segmentation phase, the input image was partitioned into three distinct color channels. Color features are then extracted individually from the resulting segmented images. In this study, the extracted color features can be considered as color-based texture features, as they are derived from first-order statistics calculated from the colored images themselves, rather than the conventional grayscale images.
Conclusion
The classification of plant species based on their leaf images using computer vision algorithms presents a significant challenge in categorizing the images into distinct groups. The similarity between plant species both within and across classes, along with the potential presence of complex backgrounds and variations in parameters such as illumination and color, further compounds the difficulty. Hence, there is a need for effective tools and solutions to analyze and interpret patterns in leaf images to achieve meaningful results. This project proposes an automated plant identification system that leverages deep convolutional neural networks (CNN) to achieve higher accuracy in identifying plant species based on their leaves. The system comprises three key steps: image pre-processing, feature extraction, and recognition. The proposed CNN classifier utilizes hidden layers such as convolutional layers, max pooling layers, dropout layers, and fully connected layers to learn and extract features from the leaf images. The results obtained demonstrate a slight improvement over previous works, achieving an accuracy of 93.75% in classification. In future development and enhancement of the automated system for medicinal plant classification. One area of focus could be expanding the dataset to include a broader range of medicinal plant species, allowing for a more comprehensive classification. Additionally, incorporating additional features such as textual information and chemical composition analysis could further enhance the system\'s accuracy and provide more detailed recommendations.
References
[1] S. Mahajan, A. Raina, X.-Z. Gao, and A. K. Pandit, ``Plant recognition using morphological feature extraction and transfer learning overSVMand AdaBoost,\'\' Symmetry, vol. 13, no. 2, p. 356, Feb. 2021. [2] Beikmohammadi, K. Faez, and A. Motallebi, ``SWP-leaf NET: A novel multistage approach for plant leaf identification based on deep learning,\'\' 2020, arXiv:2009.05139. [Online]. Available: http://arxiv.org/abs/2009.05139. [3] Muneer and S. M. Fati, ``Efficient and automated herbs classification approach based on shape and texture features using deep learning,\'\' IEEE Access, vol. 8, pp. 196747-196764, 2020. [4] J. W. Lee and Y. C. Yoon, ``Fine-grained plant identification using wide and deep learning model 1,\'\' in Proc. Int. Conf. Platform Technol. Service (PlatCon), Jan. 2020, pp. 1-5. [5] P. Jasitha, M. R. Dileep, and M. Divya, ``Venation based plant leaves classification using GoogLeNet and VGG,\'\' in Proc. 4th Int. Conf. Recent Trends Electron., Inf., Commun. Technol. (RTEICT), May 2020, pp. 715-719. [6] A. Kaya, A. S. Keceli, C. Catal, H. Y. Yalic, H. Temucin, and B. Tekinerdogan, ``Analysis of transfer learning for deep neural network based plant classification models,\'\' Comput. Electron. Agricult., vol. 158, pp. 20-29, Mar. 2021. [7] V. Bodhwani, D. P. Acharjya, and U. Bodhwani, ``Deep residual networks for plant identification,\'\' Proc. Comput. Sci., vol. 152, pp. 186-194, Jan. 2020.
Copyright
Copyright © 2023 Hemalatha ., Sreerambabu , Rajkumar , Santhosh . This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET55008
Publish Date : 2023-07-25
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

