Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- Conclusion
- References
- Copyright
Brain Tumor Detection and Classification using Deep Learning
Authors: Fatima Banoo, Heena Arora
DOI Link: https://doi.org/10.22214/ijraset.2023.55754
Certificate: View Certificate
Abstract
This paper explores the application of convolutional neural networks (CNNs) in diagnosing brain tumors, focusing on children and the elderly who are most vulnerable to this type of cancer. Brain tumors pose significant health risks due to uncontrolled cell growth within the skull, making accurate classification challenging. The study utilizes CNNs, a popular machine learning approach, to classify 3260 brain magnetic resonance images into glioma, meningioma, pituitary, and no tumor categories. Three algorithms, CNN, VGG, and Densenet, are employed for classification. Early detection is vital for effective treatment, and computer-aided diagnostics show promise in assisting clinicians. This work provides an overview of traditional computer-aided tumor diagnosis and highlights the advantages of using CNNs. It discusses image segmentation, classification techniques based on CNN, Densenet, and VGG16, and serves as a guide for future tumor detection research. Data augmentation and min-max normalization enhance tumor cell contrast. The dense CNN model accurately classifies a small image database, and Densenet demonstrates remarkable performance. Experimental results show CNN achieving 45% accuracy on testing data, VGG achieving 51%, and Densenet achieving an impressive 86.49% during training, with high precision and a good F1 score. Overall, this paper emphasizes the potential of CNNs for brain tumor diagnosis and contributes to the development of computer-aided diagnostic systems. It advances the field of medical image analysis and paves the way for further research in tumor detection.
Introduction
I. Introduction
Benign tumors in the human body can arise from factors such as organ dysfunction, local pressure, and congestion. These tumors often develop slowly and generally do not pose an immediate threat to the patient's life. However, as the tumor grows, it can exert pressure on the surrounding tissues, leading to various symptoms. For instance, compression of the chest may result in difficulties in breathing, chest discomfort, coughing, tightness, as well as abdominal symptoms like loss of appetite, nausea, abdominal pain, distension, and vomiting. In some cases, invasive surgeries may relieve these symptoms, but there is also a risk of malignant lesions forming, which can give rise to endocrine disorders, bleeding, and infections.
Unlike benign tumors, malignant tumors have a detrimental impact on the body. They exhibit rapid growth and immature differentiation, causing disruption to the normal function of organs by compromising their structure and proper functioning. This can greatly affect the patient's overall quality of life, potentially leading to symptoms such as coughing, hemoptysis (coughing up blood), or even impaired cardiac function. Advanced stages of cancer can result in emaciation, weakness, anemia, or systemic failure. Malignant tumors have the ability to grow and spread, leading to the development of additional systemic syndromes and an increased susceptibility to infections. It is crucial to differentiate between benign and malignant tumors as their behavior and potential consequences vary significantly. Malignant tumors, with their aggressive growth and metastatic capabilities, pose a greater threat to the patient's health and require prompt and appropriate medical intervention.
A. Methods of Tumor Diagnosis
Early detection of malignant tumors is crucial as they often do not exhibit noticeable symptoms in their initial stages. Recognizing the warning signs associated with different types of tumors can aid in their early detection and prompt initiation of treatment, leading to improved chances of successful outcomes. Thorough examinations are conducted when a tumor is suspected to obtain a comprehensive and unbiased assessment, enabling immediate treatment and enhancing the potential for a cure.
In this study, researchers utilize three fully automated models, namely CNN, VGG, and Densenet, to accurately and swiftly classify brain tumors. Despite the inherent challenges and uncertainties in brain tumor categorization, the objective is to develop autonomous CNN models with min-max normalization using publicly available datasets. The newly developed Densenet CNN architecture holds promise as a valuable decision-making tool in brain tumor diagnostic tests.
To enhance accuracy, the study proposes the utilization of a robust Densenet and VGG network for categorizing three different types of brain tumors. The focus lies on data augmentation through min-max normalization combined with denseNet to improve training speed and network depth. The inclusion of deep layers of separable convolution reduces parameters and processing requirements. Furthermore, the addition of dense blocks to the Densenet model facilitates the segmentation of brain tumors. This approach enables denseNet to achieve high classification accuracy by gathering detailed image data and generating dense segmentation masks for the classification of brain tumors using T1-weighted contrast-enhanced magnetic resonance imaging. The network's performance is evaluated through pre-processing, augmentation, and classification techniques.
The study presents an innovative deep convolutional neural network-based dense depth classifier, which outperforms existing deep learning techniques in terms of classification accuracy. The recommended methodology demonstrates excellent performance even with limited training samples, as evident from the confusion matrix. Dropout layers effectively address the problem of overfitting and contribute to reduced classification errors. Overall, the proposed approach offers promising results and contributes to the advancement of tumor classification using deep learning techniques in the field of medical imaging analysis.
II. Literature review
Hashemzehi et al. [4] conducted a study to detect brain cancers from CT Scan images using a hybrid model combining CNN and NADE. They utilized 3064 T1-weighted contrast-enhanced images and achieved a high accuracy rate of 96 percent in identifying three distinct types of brain cancers.
In another study by Diaz-Pernas et al. [5], a fully automated algorithm for brain tumor segmentation and classification based on CT scans of meningioma, glioma, and pituitary tumors was presented. They employed CNN to incorporate a multi-scale approach inspired by human functioning. Their algorithm achieved an impressive accuracy of 97 percent using a collection of 3064 slices from 233 patients.
Sultan et al. [6] employed a CNN architecture consisting of 16 convolution layers, pooling and normalization layers, and a dropout layer before the fully connected layer. They achieved a high accuracy rate of 96 percent by training the model on 68 percent of the images and validating and testing on the remaining images.
III. METHODOLOGY
In this paper, the authors conducted their research on a dataset consisting of 3260 brain magnetic resonance (MR) images, which included different types of brain tumors. The dataset was collected from Kaggle.com and comprised 3064 T1-weighted contrast-enhanced MR images. The three main types of brain tumors in the dataset were meningioma (708 images), glioma (1426 images), and pituitary tumor (930 images). The images were acquired from 233 patients and covered three imaging planes: sagittal (1025 images), axial (994 images), and coronal (1045 images).To train and evaluate their model, the authors divided the dataset into three parts: training, validation, and testing. This division allows for assessing the model's performance on unseen data. The authors applied min-max normalization and data augmentation techniques to enhance the quality and diversity of the training data.
The suggested model consists of different stages, which are visualized in Figure 1 (not provided here). The specific architecture and details of the model are not mentioned in this excerpt, but it can be inferred that the model incorporates various components or layers to process and analyze the MR images for tumor classification. Overall, we utilized a large dataset of brain MR images containing different types of tumors to develop and evaluate their model. The dataset was divided into training, validation, and testing sets, and the model architecture was structured into distinct stages..
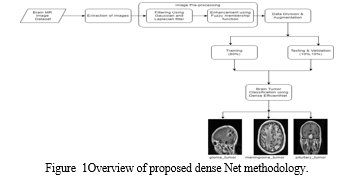
V. mage Pre-Processing
The brain tumor images in this study suffer from poor quality, including noise and low illumination. To address this issue, the proposed method employs data normalization and a specific technique called min-max normalization to enhance the brightness of the images. The authors apply Gaussian and Laplacian filters to further improve the image quality. Initially, the authors add Gaussian blur to the original images. Then, they subtract the blurred image from the original by incorporating a weighted portion of the mask, resulting in a de-blurred image. Additionally, a Laplacian filter with a kernel size of 3 × 3 is utilized to smooth the images. These steps are illustrated in Figure 2.
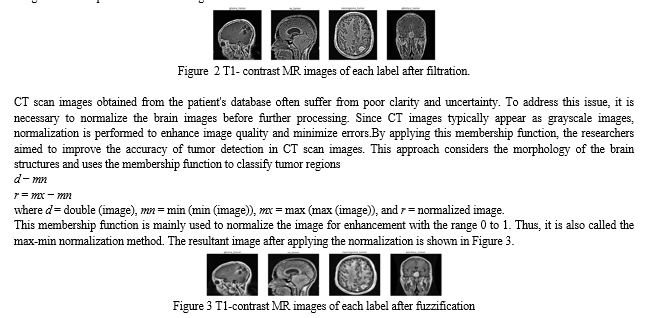
C. Data Division and Augmentation
The deep neural network needs large datasets for better results but our dataset is limited. Our dataset contains 3260 brain images, further divided into 80% for training, which remains for testing and validation purposes. So, data augmentation is needed to change in the minor. The authors have applied rotation, width-shift, height-shift, and the zoom—range for the data requirement. They augmented the original data 21 times for better training. This will enhance the amount of training data, allowing the model to learn more effectively. This may assist in increasing the quantity of relevant data. It contributes to the reduction of overfitting and enhances generalization. Data augmentation (DA) is the process of creating additional samples to supplement an existing dataset via transformation. Dropout through augmentation, practical solutions such as dropout regularization, and batch normalization are performed on the original dataset. By data warping or oversampling, this augmentation exaggerated the size of the training dataset.
D. Dense Net Model
A novel dense CNN model is presented in this article, which is a mix of pre-trained EfficientNetB0 with dense layers. EfficientB0 has 230 layers and 7 MBConv blocks [22,23]. It features a thick block structure consisting of four tightly linked layers with a development rate of 4. Each layer in this structure uses the output feature maps of the preceding levels as the input feature maps. The dense block concept is composed of convolution layers of the same size as the input feature maps in EfficientNet. Dense block takes advantage of the preceding convolution layers’ output feature maps to generate more feature maps with fewer convolution kernels. This CNN model retrieved 150 × 150 enhanced CT scan image data. The dense EfficientNet network has an alternate dense and drop-out layer. A dense layer is the basic layer which feeds all outputs from the previous layer to all its neurons, each neuron providing one output to the next layer. The drop-out layer is used to reduce the capacity or thin the network during training and avoids the overfitting. We begin by adding a pooling layer, followed by four dense layers and three drop-out layers to ensure the model runs smoothly. The numbers of neurons in the dense units are 720, 360, 360, and 180, respectively. The drop-out values are 0.25, 0.25, and 0.5, respectively.
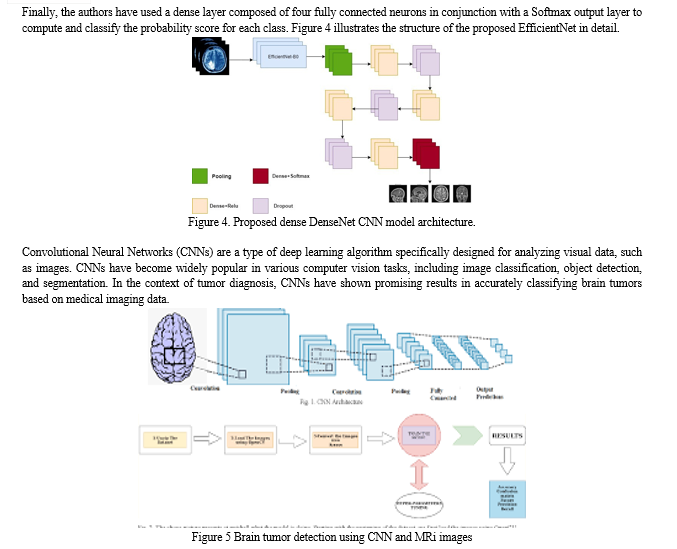
The implementation of CNNs for tumor classification involves several key steps. Convolutional layers extract local features by applying learnable filters, detecting patterns, edges, and textures at different spatial scales. Pooling layers then downsample the feature maps, aggregating neighboring values to extract relevant features while reducing computational complexity. Activation functions introduce non-linearities, allowing the CNN to learn complex relationships between input data and tumor class labels. Fully connected layers act as classifiers, using learned features to make predictions, followed by a softmax activation function for probability scores.
During training and backpropagation, the CNN adjusts its parameters using optimization algorithms to minimize a loss function. Labeled training data associates input images with tumor class labels, enabling the network to learn and improve its performance iteratively. Transfer learning is another approach, utilizing pre-trained CNN models and fine-tuning them on the tumor dataset. This can enhance training speed and performance, especially with limited tumor data available. By implementing a CNN for tumor classification, relevant features can be automatically extracted from medical images, enabling predictions about the presence and type of brain tumors. This involves defining the architecture, training the model, fine-tuning hyperparameters, and evaluating performance metrics to assess the model's classification capabilities.
E. VGG
VGG (Visual Geometry Group) is a deep convolutional neural network architecture that has demonstrated excellent performance in image classification tasks. It was developed by the Visual Geometry Group at the University of Oxford and has been widely adopted in computer vision applications, including tumor classification in medical imaging.
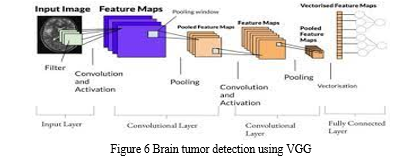
The implementation of VGG for tumor classification involves key steps. The architecture consists of convolutional layers that extract features, capturing patterns at different scales. Max pooling layers downsample the feature maps, preserving important features. Fully connected layers act as a classifier, making predictions using the learned features and a softmax activation function. Training includes updating parameters through optimization algorithms and backpropagation.
Transfer learning can be employed by utilizing pre-trained VGG models and fine-tuning them on the tumor dataset. This accelerates training and improves performance, especially with limited data. The implementation process includes defining the architecture, training the model, fine-tuning hyperparameters, and evaluating performance metrics.
VGG's deep architecture enables it to capture intricate details and learn complex representations. By implementing VGG for tumor classification, the model can effectively analyze medical images and provide accurate predictions about the presence and type of brain tumors.
IV. Simulation and results
Firstly the necessary libraries are and setting up the directories for training, validation, and testing data. It also includes a function to randomly view images from the dataset. The imported libraries include numpy, pandas, tensorflow, matplotlib, and keras, among others. These libraries provide various functionalities for data manipulation, deep learning, and visualization.
The code sets the directory paths for the training, validation, and testing data. The train_dir, val_dir, and test_dir variables specify the respective directories' paths.
The class_names variable is created to store the names of the different tumor classes present in the dataset.
The view_random_image function takes a target directory and class as input and randomly selects an image from that class's folder. It then displays the image using matplotlib.
Lastly, the code uses a loop to display 12 randomly selected images from the dataset, with their corresponding class names.
Overall, this code segment sets up the necessary libraries, directory paths, and functions for further data exploration and image visualization.
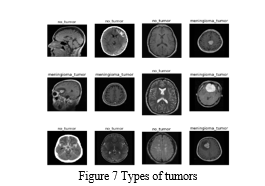
The random seed is set to ensure replicable results.
The IMAGE_SIZE variable is set to (224, 224), indicating the desired size of the images in the dataset.
The BATCH_SIZE variable specifies the number of images in each batch during training. The VAL_SPLIT variable determines the proportion of data that will be used for validation (20% in this case).
The tf.keras.preprocessing.image_ dataset_from_ directory function is used to create three datasets: train_data, valid_data, and test_data. These datasets are created by loading images from their respective directories (train_dir, val_dir, and test_dir). The function also performs preprocessing operations such as resizing the images to IMAGE_SIZE and converting the labels to categorical format.
Data augmentation is applied to the train_data dataset using the data_augmentation sequential model. This includes random horizontal and vertical flips, random rotations, and random zooms. The augmented data is then mapped to the train_data dataset. The code segment prepares the datasets for training, validation, and testing by resizing the images, performing data augmentation on the training data, and organizing the data into batches. These steps are essential for training a deep learning model on the provided image data.
F. Training Model
The callbacks module from tensorflow.keras is imported. An EarlyStopping callback is created with the following parameters:
min_delta: The minimum change in the monitored quantity to qualify as an improvement.
patience: The number of epochs with no improvement after which training will be stopped.
restore_best_weights: Whether to restore the weights from the epoch with the best value of the monitored quantity.
The VGG16 model is loaded from tensorflow.keras.applications with pre-trained weights from ImageNet. The include_top parameter is set to False to exclude the fully connected layers of the VGG16 model. The input_shape parameter is set to (224, 224, 3) to match the input size of the images.
The base_model is set to be non-trainable by setting base_model.trainable to False. This means that only the top layers added to the model will be trained. The model_Inc sequential model is created, starting with the resize_and_rescale sequential model, followed by the base_model. The GlobalAveragePooling2D layer is added to flatten the features, and a dropout layer with a dropout rate of 0.5 is added for regularization. Then, two fully connected (Dense) layers with 64 and 16 units, respectively, and ReLU activation are added. Finally, a Dense layer with 4 units and softmax activation is added for multi-class classification.The model is compiled with a categorical_crossentropy loss function, an Adam optimizer with a learning rate of 0.001, and accuracy as the evaluation metric.
The code segment creates a VGG16-based model for tumor classification and configures it for training. The EarlyStopping callback is set up to monitor the validation loss and stop training if the loss does not improve for a certain number of epochs.
The code provided trains the model_Incep model on the train_data with validation on the valid_data for a specified number of epochs. The training progress is displayed for each epoch.
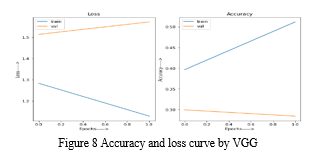
After training, the history of training and validation loss and accuracy is stored in a dataframe df. This dataframe is then used to plot two subplots: one for the loss values (training and validation) and one for the accuracy values (training and validation).
The resulting plot shows the trend of the loss and accuracy values over the epochs. The first subplot displays the loss values, while the second subplot displays the accuracy values. The x-axis represents the number of epochs, and the y-axis represents the corresponding loss or accuracy values. The plot helps visualize the training progress and the performance of the model on the training and validation data.
It is important to note that in the code snippet provided, the number of epochs specified is 2, but the initial value of EPOCHS is set to 1.
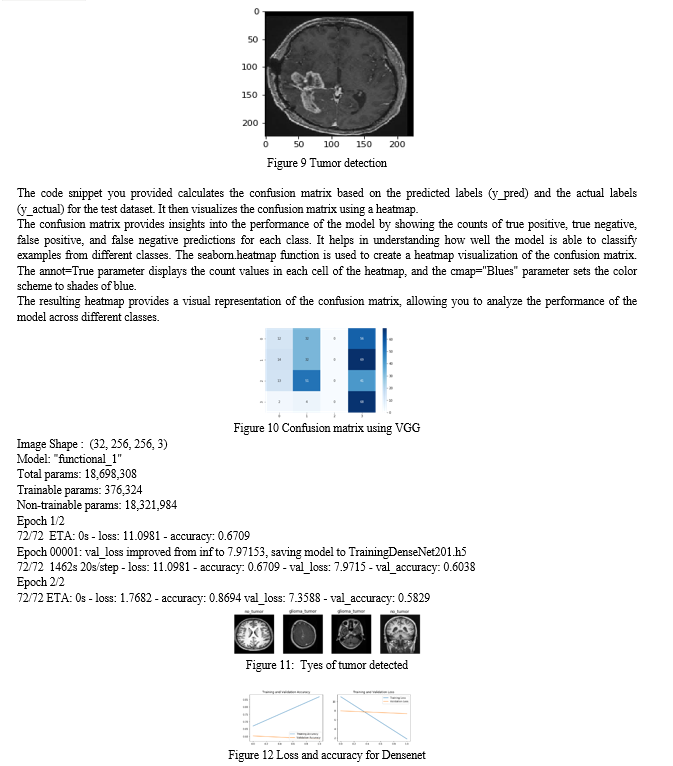

Conclusion
This paper focused on the application of Convolutional Neural Networks (CNNs) for the diagnosis of brain tumors, with a particular emphasis on children and the elderly who are most susceptible to this form of cancer. The challenges associated with categorizing tumor cells accurately were addressed, considering their diverse nature. CNNs, a widely used machine learning approach for image recognition and visual learning, were explored as a potential solution. The research aimed to classify 3260 T1-weighted contrast-enhanced brain magnetic resonance images into four groups: glioma, meningioma, pituitary, and no tumor. Three machine learning algorithms, namely CNN, VGG, and Densenet, were employed to accomplish this classification task. The paper provided an overview of traditional computer-aided tumor diagnostic approaches and emphasized the advantages of using CNNs for tumor diagnosis. It discussed the segmentation and classification of tumor images, along with the diagnosis techniques based on CNN, Densenet, and VGG16. The research serves as a guide for the development of CNN-based computer-aided systems for future tumor detection research. To enhance the contrast of tumor cells, the authors utilized data augmentation in conjunction with min-max normalization. The dense CNN model demonstrated its effectiveness in accurately classifying a small database of images. Additionally, Densenet exhibited remarkable overall performance. The experimental results indicated that the suggested CNN achieved an accuracy of 45% on testing data, while VGG attained 51% accuracy. Notably, Densenet achieved an impressive accuracy of 86.49% during training, demonstrating its superior performance. Both Densenet and VGG also exhibited high precision and a good F1 score, further highlighting their potential in tumor diagnosis. Overall, this paper emphasized the potential of CNNs in brain tumor diagnosis and contributed insights into the development of computer-aided diagnostic systems. The findings add value to the field of medical image analysis and lay the groundwork for future advancements in tumor detection research. The utilization of CNNs and other machine learning algorithms holds promise in improving the accuracy and efficiency of brain tumor diagnosis, ultimately benefiting patients and clinicians in their fight against cancer.
References
[1] Pradhan, A.; Mishra, D.; Das, K.; Panda, G.; Kumar, S.; Zymbler, M. On the Classification of MR Images Using “ELM-SSA” Coated Hybrid Model. Mathematics 2021, 9, 2095. [CrossRef] [2] Reddy, A.V.N.; Krishna, C.P.; Mallick, P.K.; Satapathy, S.K.; Tiwari, P.; Zymbler, M.; Kumar, S. Analyzing MRI scans to detect glioblastoma tu-mor using hybrid deep belief networks. J. Big Data 2020, 7, 35. [3] Nayak, D.R.; Padhy, N.; Mallick, P.K.; Bagal, D.K.; Kumar, S. Brain Tumour Classification Using Noble Deep Learning Approach with Parametric Optimization through Metaheuristics Approaches. Computers 2022. [4] Mansour, R.F.; Escorcia-Gutierrez, J.; Gamarra, M.; Díaz, V.G.; Gupta, D.; Kumar, S. Ar-tificial intelligence with big data analytics-based brain intracranial hemorrhage e-diagnosis us-ing CT images. Neural Comput. Appl. 2021. [CrossRef] [5] Rehman, A.; Naz, S.; Razzak, M.I.; Akram, F.; Imran, M.A. Deep learning-based framework for automatic brain tumorsclassification using transfer learning. Circuits Syst. Signal Processing 2020, 39, 757–775. [CrossRef] [6] Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [7] Ozyurt, F.; Sert, E.; Avci, D. An expert system for brain tumor detection, Fuzzy C-means with super-resolution and convolutional neural network with extreme learning machine. Med. Hypotheses 2020, 134, 109433. [8] Hu, M.; Zhong, Y.; Xie, S.; Lv, H.; Lv, Z. Fuzzy System Based Medical Image Processing for Brain Disease Prediction. Front. Neurosci. 2021, 15, 714318. [CrossRef] [PubMed] [9] Maqsood, S.; Damasevicius, R.; Shah, F.M. An Efficient Approach for the Detection of Brain Tumor Using Fuzzy Logic and U-Net CNN Classification. In Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2021; Volume 12953. [10] Ragupathy, B.; Karunakaran, M. A fuzzy logic-based meningioma tumor detection in magnetic resonance brain images using CANFIS and U-Net CNN classification. Int. J. Imaging Syst. Technol. 2021, 31, 379–390. [11] Cheng, J.; Huang, W.; Cao, S.; Yang, R.; Yang, W.; Yun, Z.; Wang, Z.; Feng, Q. Correction, enhanced performance of brain tumor classification via tumor region augmentation and partition. PLoS ONE 2015, 10, e0144479. [CrossRef] [12] Badža, M.M.; Barjaktarovic´, M.C. Classification of brain tumors from MRI images using a convolutional neural network.? Appl. Sci. 2020, 10, 1999. [CrossRef] [13] Mzoughi, H.; Njeh, I.; Wali, A.; Slima, M.B.; BenHamida, A.; Mhiri, C.; Mahfoudhe, K.B. Deep multi-scale 3D convolutional neural network (CNN) for MRI gliomas brain tumor classification. J. Digit. Imaging 2020, 33, 903–915. [CrossRef] [PubMed]
Copyright
Copyright © 2023 Fatima Banoo, Heena Arora. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET55754
Publish Date : 2023-09-16
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

