Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- Conclusion
- References
- Copyright
A Multi-View Learning based Clustering Method for Health Care System
Authors: Neha Garg, Sunidhi Shrivastava
DOI Link: https://doi.org/10.22214/ijraset.2022.43243
Certificate: View Certificate
Abstract
Patients in hospitals have faced complications due to measurement errors, missing data, privacy issues etc. in electronic medical records. However, these medical records from heterogeneous sources have both structured and unstructured data. In particular, unstructured clinical data is valuable source of information including patient’s records of pathology data, radiology findings, medication order etc. However, to scrutinize, construe and presentation of this unstructured and high dimensional data is one of the significant modeling challenge that clinical support system has faced from many years before. Therefore, there is a need of some standard technique to locate both subjective and objective guesstimates of patient’s condition. Our endowments in this paper are twofold. First, present a multi-view learning technique, i.e. Collective Matrix Factorization to combine the extracted features from multiple views and gives a low dimensional representation of combined clinical data. Second, proposed a Genetic-K-means based clustering algorithm based on Collective Matrix Factorization for heterogeneous clinical records. It has been observed by the experiments that proposed method gives more accurate clustering results than existing method
Introduction
- INTRODUCTION
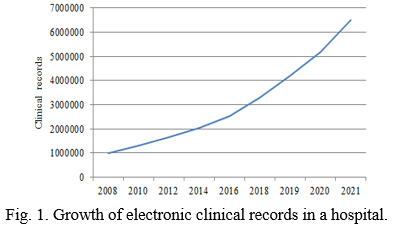
Nowadays, a gigantic amount of unstructured data is generated in health care system. These data contains patient records of various tests, diagnosis and treatments. In the past years, it is estimated that the growth of the unstructured data is about 80% and only 20% of structured data. However, it is arduous to scrutinize this unstructured data of electronic clinical records and find the critical information to improve the patient healthcare process. Figure 1 portrays the explosion of electronic health records over the past 15 years. This volume of data contains various medical records of diagnosis data, pathology data, radiology data, nursing data, vital data etc., each have numerous different structured and unstructured documents that are typically arduous to extract meaningful knowledge due to the enormous size of electronic medical records in hospitals. However, the problems are associated with unstructured and high dimensional data such as radiological findings, diagnosis data etc. that also contains meaningful information about the patients. In addition, there are various complications that patients have faced in hospitals due to measurement errors in data, missing data, privacy issues etc. [1] Therefore, to scrutinize and construe this data is one of the important challenges that clinical support system has faced from many years before [2]. There is a need of some standard tools to locate both subjective and objective approximations of patient’s condition.
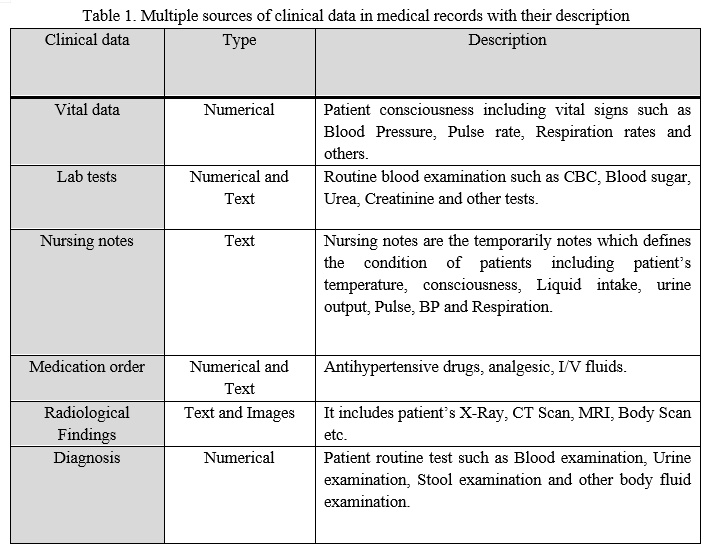
When a patient enters in a hospital they have passed many stages and at each stage, clinical data of a patient is recorded either digitally in electronic records or on paper. These clinical data includes nursing notes, vital data, diagnoses, pathology data, radiology images etc. Table 1 illustrates the description of these various sources of patient data available in hospitals records
II. BACKGROUND HISTORY
There have been many efforts for exploiting the heterogeneity of clinical records in health care [3].
A. Disease Progression Model
One application for health care is disease progression model. Wang et al. [4] works on this model using incomplete and heterogeneous clinical records. Cook et al. [5] presents the novel applications which integrate disease progression model with cost effectiveness analysis and genomic data analysis. Cohen et al. [6] uses bioinformatics cluster analysis for identification of multifaceted metabolic states of patients.
B. Time series model
Other application for health care is time series model. Bui et al. [7] has review different types of forecasting model for medical purposes using time series based methods which helps the researcher to determine suitable forecasting models. Mao et al. [8] develops an integrated mining approach to provide worsening caveats for patients in RDS and ICU. Yang et al. [9] develop a data mining time series method to forecast days in hospitals for scrutinizing health insurance claims. Durichen et al. [10] presents multi-task GPs for multiple correlated multivariate physiological time series. Batal et al. [11] presents a framework for classifying multivariate data based on temporal pattern.
In this article, we scrutinizes the complications that arising in the critical care and effectively handle by the expansion of text mining tools. We also define the multi view technique to effectively combine the numerical as well as text based features that are obtained from the vital data with other clinical data for extrapolative modeling. We present an effective text mining model for clustering of these extracted features on the basis of hybrid clustering algorithm.
The further section of this article is systematized as follows: Methods of text preprocessing and feature extraction have been described in Section 3. Section 4 defines the proposed method for clustering clinical records. Section 5 presents the performance evaluation of clustering algorithms. Section 6 concludes the paper.
III. TEXT MINING IN HEALTH CARE
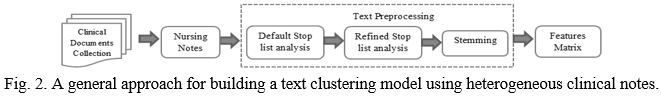
Text mining provides an opportunity to discover knowledge from these textual medical data archives. In the field of medical care, patient information is distributed in various forms such as medical tests, pathology tests, radiology images etc. This information can be effectively used to develop intelligent models to improve health care systems. Figure 2 refers to the general approach of developing intelligent models for text analysis.
A. Text Preprocessing and Feature Extraction for an extrapolative system
The text mining process starts with collecting text documents that are to be studied. The next step that have done is text preprocessing. The text preprocessing is applying on the patients records that have suffering from various diseases to remove the irrelevant words from the records. The key idea of text preprocessing is that realizing the importance of words in documents. To perform preprocessing, default stop words removal list is being used which contains terms that we have ignored in the patient records that don’t gives much value eg. frequently occurring terms such as “the, on, he, she, of” etc. are removed from the dataset. We also have create medical related stop words list by removing the terms that are medically irrelevant (such as “prepped”, “position”, “passed”, “monitored” etc.) and kept only the terms including the symptoms of diseases with their medications. The importance of same word has differed when it comes under the different records. The preprocessing result gives the entire features vector that occurs in the records along with their importance to each word represented as frequencies.
B. Feature Extraction using Multi-view learning approach
In the field of medical care, patient information is disseminated in numerous forms such as nursing notes, pathology tests, radiology images etc. Extracting features from these heterogeneous data presents a different and multiple views for the same subject. In Multi-view learning approach, aim to concatenate all the features from each data and represented them as single dataset. There are several studies [12] that have exemplified the benefits of multi view data over simple concatenation. Most of these studies are based on Canonical Correlation Analysis (CCA) [13] which assesses the linear relationships between multidimensional variables.
Klami, A. et al. [14] presents a novel multi view learning based method which allows each of the views have separate low rank structure that is independent of other views as well as it supports binary, continuous and count observations that is efficient for sparse matrices which involve missing data. Later, in 2016, Huddar et al. [15] proposed a new preprocessing technique for clinical database that has been used in classifying model for identifying patient at risk. They also present a multi view approach for modeling heterogeneous clinical notes. Therefore, this method gives several benefits for incorporate in clinical datasets that have multiple views including features of nursing data, blood tests, radiology images etc. The brief illustration of Collective Matrix Factorization (CMF) approach of [14, 15] is defined in figure 3.
A matrix is defined as the relationship between the set of entities. Entity ei signifies the entity set for the rows and columns of corresponding matrix. As shown in figure, entity e1 represents the set of patients, e2 represents the set of variables to nursing notes and X1 represents the relationship between e1 and e2 i.e. it represents the nursing notes for the set of patients.
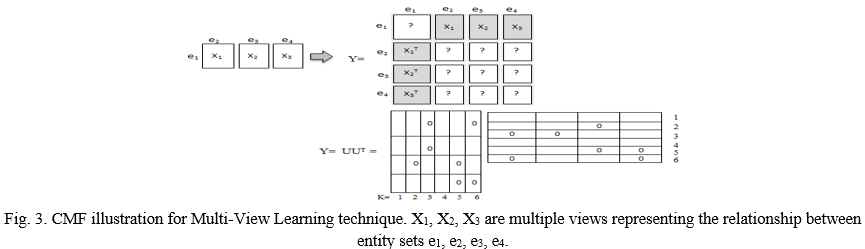
In the same way, X2 represents the relationship between e1 and e3 in which e3 represents the set of variables related to lab tests and so on.
Given M matrices Xm = [xij (m)] defines the E set of entities having cardinality de. Let rm and cm represents the entity sets equivalent to rows and columns of mth matrix. The component equivalent to row i and column j of the mth matrix can be composed as [14]:
xij (m) = k=1ku ik(rm) ujk (cm) + bi (m, r) + bj (m, c) + ?ij (m) (1)
Where Ue = [uik(e)] € R de*k defines the low rank matrix which is related to e (entity set), ?ij (m) delimits the element-wise independent noise and bi (m,r) and bj (m,c) defines the bias terms for mth matrix. The same model is attained by collecting all matrices into a large symmetric matrix Y of dimension d=e=1Ed e. As shown in matrix, some spaces are left blank, because there is no relationship between the entities.
The CMF model is:
Y=UUT + ? where U € R d*k denotes the concatenation of all Ue matrices. The block corresponds to entity e1 in matrix U represents the joint operation of all matrices for e1 entity set. This article relates to patient that used as combined representation of patients records that shared across all views.
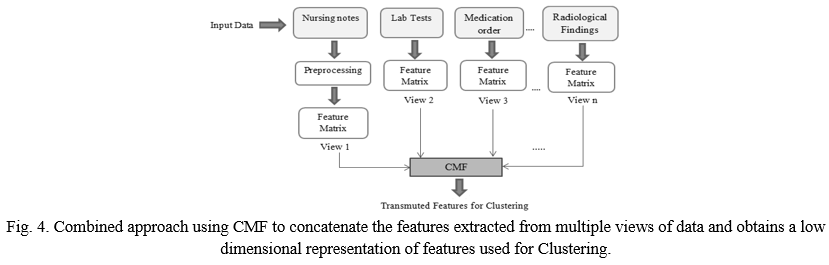
C. The Combined Approach
Figure 4 depicts the combined approach for obtaining a low dimensional representation of features. Features are extracted from patient’s nursing notes, lab tests, medication order, radiological findings, diagnosis etc. These extracted features are represented as multiple views that are combined by CMF which obtain transformed features for clustering. As shown in figure 3, entity e1 represents the set of patients, e2 represents the text feature matrix X1 from view 1. In the same way, e3 represents the feature matrix X2 from view 2 and so on. Thus, entities e2, e3, e4…..represents the feature matrices from different views and matrixes X1 has text features while matrixes X2, X3, X4….., have statistical feature from patient’s lab tests, medication order, radiological findings etc. This different view of matrices is given as input to CMF which gives the output matrix U. As shown in figure, matrix U has blocks corresponding to entities e1, e2….., and these block has a k dimensional representation of patients (k is empirically chosen) that gives input as transformed feature for clustering.
IV. THE PROPOSED ALGORITHM
The text mining process starts with collecting patient’s text records such as nursing notes, medication order, lab tests etc. then preprocessing is done to extract the features. There are numerous methods of preprocessing such as stop words removal, stemming etc. which gives the interesting patterns [16]. The next step is document representation which gives the feature matrix using TF-IDF [17]. These extracted feature matrixes are represented as multiple views that are combined by CMF which gives transformed features for clustering.
The proposed method is the combination of Genetic and K-means algorithm [24] which takes the CMF feature matrix as input. The Genetic algorithm (GA) is starts with defining initial population of chromosomes [18]. In this article, GA is use for defining initial cluster centroids of K-means algorithm for improving the accuracy of method. Therefore, a chromosome is a vector of k cluster centroids of initial clusters that is initialized to k randomly chosen rows [19] from CMF generated feature matrix.
Chromosome chr = (center1, center2,….., centerk) (2)
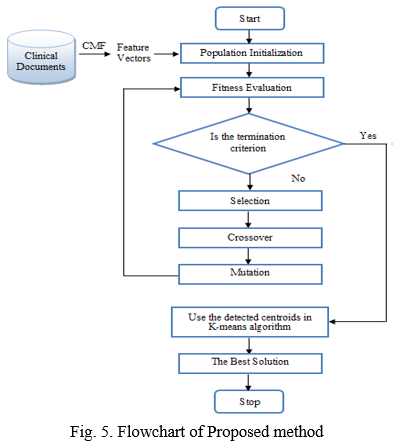
In GA [20], the fitness function is use to be 1/DB in which DB delineates the well-known Davies-Bouldin index [21, 22]. The proposed method employs the single point crossover and the mutation is performed by using Gaussian distribution [23].
V. PERFORMANCE EVALUATION
Three medical disease dataset were used during evaluation of results. A brief description of datasets [27] is given below:
- BUPA Liver Disorder Dataset-This dataset contains`345 instances and 6 numeric attributes in which first 5 attributes are all blood tests which are thought to be sensitive to liver disorders. The 6th attribute corresponds to the number of alcoholic beverages drunk per day.
- Thyroid disease Dataset- This dataset having separated training and testing instances with 3 classes. The training set has 3772 instances and the testing set has 3428 instances.
- Statlog Heart Dataset- This dataset have 270 instances with 13 attributes in which 6 real attributes, 1 ordered, 3 binary and 3 nominal.
The performance evaluation of algorithms is based on two measures: F-measure and purity [25, 26]. Figure 6 and Figure 7 gives the comparative results of existing and proposed algorithm with respect to F-measure and purity measure respectively. From the results, it could be seen that proposed method improves accuracy of clusters as compared to the existing method.
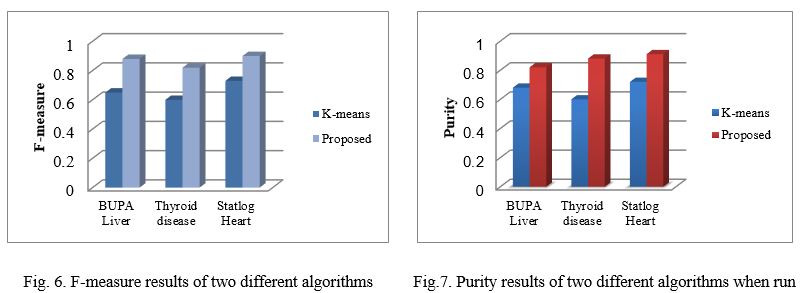
Figure 8 shows the execution time of the two different methods with respect to clusters size. From the results, it could be seen that proposed method takes more time in execution of algorithm as compared to the existing method.
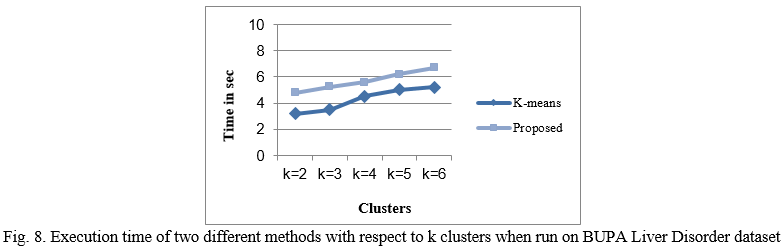
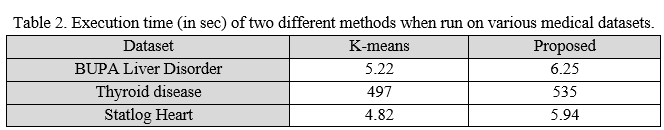
Table 2 depicts the execution time of the two algorithms when run on three different datasets. As it can be seen, the existing algorithm runs faster than the enhanced method. The possible reason for slowness of proposed method includes more calculations, improves the quality of clusters which gives more accurate results. Medical field is more concerned about the accuracy rather than time. Therefore, proposed method is considered as best algorithm for clustering of clinical records as compared to existing algorithm
Vi. FUTURE SCOPE
This proposed method can be applied on other health care datasets to improve the medical results and also use other data mining algorithms to perform clustering.
Conclusion
In the field of medical care, patient information is distributed in various forms such as nursing notes, vital data, lab tests etc. Extracting features from these unstructured data is a significant challenge in health care system. In this paper, we present a Collective Matrix Factorization (CMF), a multi view technique to combine the extracted features from multiple views and provide a low dimensional representation of combined data. These combined features are given as input to the proposed clustering method, a combination of Genetic and K-means algorithm. An experimental result proves improved accuracy of proposed method over existing method
References
[1] Johnson, E. W., Ghassemi, M. M., Nemati, S., Niehaus, K. E., Clifton, D. A. and Clifford, G. D. ( 2016). Machine learning and decision support in critical care, Proc. IEEE, 104(2):444-466. [2] Desai, S. V., Law, T. J. and Needham, D. M. (2011). Long-term complications of critical care, Critical Care Med, 39(2):371-379. [3] Reddy, C. K. and Aggarwal, C. C. (2015). Healthcare Data Analytics, 36, Boca Raton, FL, USA, CRC Press. [4] Wang, X., Sontag, D. and Wang, F. (2014). Unsupervised learning of disease progression models, in Proc. 20th ACM SIGKDD Int. Conf. Knowl. Discovery Data Mining, 85-94. [5] Cook, S. F., Bies, R. R. (2016). Disease Progression Modeling: Key Concepts and Recent Developments, Curr Pharmacol Rep., 2(5): 221-230. [6] Cohen, M. J., Grossman, A. D, Morabito, D., Knudson, M. M., Butte, A. J. and Manley, G. T, (2010). Identification of complex metabolic states in critically injured patients using bioinformatic cluster analysis, Critical Care, 14(1). [7] Bui, C., Pham, N., Vo A., Tran, A., Nguyen, A., Le, T. (2018). Time Series Forecasting for Healthcare Diagnosis and Prognostics with the Focus on Cardiovascular Diseases, In: Vo Van T, Nguyen Le T, Nguyen Duc T. (eds) 6th International Conference on the Development of Biomedical Engineering in Vietnam (BME6). BME. IFMBE Proceedings, vol 63:809-818. Springer, Singapore. [8] Mao, Y.,Chen, W., Chen, Y., Lu, C., Kollef, M. and Bailey, T. (2012). An integrated data mining approach to real-time clinical monitoring and deterioration warning, In Proc. 18th ACM SIGKDD Int. Conf. Knowl. Discovery Data Mining,1140-1148. [9] Xie, Y., Schreier, G., Hoy, M., Liu, Y., Neubauer, S., Chang, D. C. (2016). Analyzing health insurance claims on different timescales to predict days in hospital, J Biomed Inform. [10] Dürichen, R., Pimentel, M A F., Clifton, L., Schweikard, A. and Clifton, D A. (2015). Multitask Gaussian processes for multivariate physiological time-series analysis, IEEE Trans. Biomed. Eng., 62(1):314-322. [11] Batal, I., Valizadegan, H., Cooper, G. F. and Hauskrecht, M. (2011). A pattern mining approach for classifying multivariate temporal data, in Proc. IEEE Int. Conf. Bioinformatics Biomed. (BIBM), 358-365. [12] Hardoon, D. R., Szedmak, S. and Shawe-Taylor, J. (2003). Canonical correlation analysis: An overview with application to learning methods, Neural Comput., 16(12):2639-2664. [13] Hotelling, H. (1936). Relations between two sets of variates, Biometrika, 28(3/4): 321-377. [14] Klami, A., Bouchard, G. and Tripathi, A. (2014) Group-sparse embeddings in collective matrix factorization, in Proc. Int. Conf. Learn. Represent. (ICLR). [15] Huddar, V, Desiraju,. B. K., Rajan, V., Bhattacharya, S., Roy, S., Reddy, C. K. (2016) Predicting Complications in Critical Care using Heterogeneous Clinical Data, Special Section on Big Data Analytics for Smart and Connected Health, IEEE Access 4, 7988-8001, 2016. [16] Han, J., Kamber, M. (2006) Data Mining: Concepts and Techniques, Morgan Kaufmann, 2nd Ed. [17] Salton,. G, Wong, A., Yang, C. S. (1975) A Vector Space Model for Automatic Indexing, In: Communications of the ACM, 18(11):613-620. [18] Holland, J. (1975) Adaptation in Natural and Artificial Systems, University of Michigan Press. [19] Garg, N., Gupta, R. K. (2018) Performance Evaluation of New Text Mining Method Based on GA and K-Means Clustering Algorithm. In: Choudhary R., Mandal J., Bhattacharyya D. (eds) Advanced Computing and Communication Technologies. Advances in Intelligent Systems and Computing, vol 562. Springer, Singapore. [20] Goldberg, D. E. (1989). Genetic Algorithms in Search. Optimization and Machine Learning, Addison Wesley Publishing Company. [21] Davies, D. L. and Bouldin, D. W. (1979) A cluster separation measure, IEEE Trans. Pattern Anal. Intell., 1(2): 224-227. [22] Bandyopadhyay, S. and Mauilk, U. (2001). Nonparametric genetic clustering: Comparison of validity indices, IEEE Trans. System Man Cybern.-Part C Applications and Reviews, 31:120-125. [23] Yao, X., Liu, Y., Lin, G. (1999). Evolutionary programming made faster. In: IEEE Transactions on Evolutionary Computation, 3(2): 82-102. [24] Bhatt, V., Dhakar, M., Chaurasia, B. K. (2016) Filtered Clustering Based on Local Outlier Factor in Data Mining, Inernational Journal of Database Theory and Application, 9(5):275-282. [25] Aliguliyev, R. M. (2009). Clustering of document collection - A weighting approach, In: Expert Systems with Applications, 36(4):7904-7916. Elsevier. [26] Abraham, A., Das, S., Konar, A. (2006). Document Clustering using Differential Evolution, In: IEEE Congress on Evolutionary Computation CEC,1784-1791. [27] Datasets,http://archive.ics.uci.edu/ml/datasets.html.
Copyright
Copyright © 2022 Neha Garg, Sunidhi Shrivastava. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET43243
Publish Date : 2022-05-25
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

