Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- Conclusion
- References
- Copyright
Genome Sequence Analysis of Lungs Cancer Protein WDR74 (WD Repeat-Containing Protein)
Authors: Navjot Kaur Virk, Uma Kumari
DOI Link: https://doi.org/10.22214/ijraset.2022.43175
Certificate: View Certificate
Abstract
There are over 200 distinct forms of cancer, and all are diagnosed and treated differently. According to the WHO Global Cancer Observatory, 19,292,789 cases new cases of cancer were diagnosed in 2020, with breast (11.7% cases), lung (11.4%), and Colorectum (10% cases) becoming the three most common. Lung cancer is one of the most often diagnosed malignancies and the biggest cause of cancer-related deaths globally, with an estimated 220 million new cases and 179 million deaths every year. It is extremely invasive, quickly spreading, and cause death in both sexes. In light of both core genetic abnormalities and therapy response, lung cancer is a highly diverse ailment. WDR74 protein predominantly controls WNT signaling pathways in lung cancer. Wnt-responsive genes such as c-myc and cyclin D1 have been linked to cell proliferation, whereas caspase 3, caspase 9, and MDR1 have been linked to chemo-resistance and death. WDR74 had regulatory impacts on these genes. In lung cancer cells, WDR74 influenced several biological processes in lung cancer cells by modulating these genes. Abnormal activation of the Wnt/-catenin signalling pathway promotes a number of cellular functions such as proliferation, cell cycle progression, aggressiveness, and chemoresistance, specifically in Lung cancer. Our study uses local alignment to establish WDR74 sequence similarity, and then uses multiple sequence alignment to compare homologous sequences. We then used software to scan the sequence for open reading frames (ORFs) to see if WDR74 had main mutations. The study\'s findings include a brief summary of the top five protein matches from well-studied reference species in the database, as well as a graphical summary and phylogenetic tree development. The study also suggested the open reading frame with the primary mutation, as well as the start and stop codon positions.
Introduction
I. INTRODUCTION
Lung cancer is still the major cause of death from cancer in both sexes in the United States [1] and throughout the world [2]. Approximately 90% of lung cancer cases are linked to smoking and the use of tobacco and other lung cancer-causing factors such as asbestos, air pollution exposures, chronic infections, and so on [3]. According to the WHO Global Cancer Observatory, 19,292,789 cases new cases of cancer were diagnosed in 2020, out of which the death rate due to lungs cancer was the greatest that was 18%. Lung cancer is classified into two main types based on the growth and spread is small-cell lung carcinomas and non-small cell lung carcinomas [4]. Small-cell lung carcinomas account for 20–25% of all lung malignancies and propagate into sub-mucosal lymphatic vessel and regional lymph nodes, whereas 85% of diagnoses were made for NSCLC and acts akin to SCLC in terms of rapid deadly spread [5]. Squamous-cell carcinomas, adenocarcinomas, and large-cell carcinomas are subtypes of non-small-cell carcinomas [6]. The most frequent kind of lung cancer among nonsmokers and women is adenocarcinoma. It account for 30–40% of lung cancers. They tend to develop toward the lung's periphery. Lung malignancies with squamous-cell carcinomas make up around 30% of all cancers, and are more commonly detected in the center portion of the chest and develop slowly. Large cell carcinoma is the least prevalent lung cancer, contributing to lung cancer, contributing to 10–15 percent of all cases. These tumors usually form on the periphery, and have a higher proclivity for spreading [7]. Many genes alteration, including EGFR, KRAS, MET, LKB1, BRAF, PIK3CA, ALK, RET, and ROS1, can cause lung cancer [8]. Likewise, the WDR74 protein i.e. WD repeat-containing protein has been found to be important in the development of lung cancer and melanoma primary tumors and metastases. WDR74 is a key regulator of embryogenesis and cell growth [9]. WDR74 acts as a 60S ribosome assembly factor that is in charge of protein synthesis in every live cell [10]. Its expression was two times greater in metastatic circulating tumor cells than in the primary tumor. WDR74 protein predominantly controls WNT signaling pathways in lung cancer cells that are activated in tumor cells. In many malignancies, abnormal activation of the Wnt/-catenin signalling pathway promotes a number of cellular functions such as proliferation, cell cycle progression, aggressiveness, and chemoresistance, specifically in Lung cancer. Overexpression of WDR74 inhibited phosphorylation-dependent degradation of β-catenin and increased its translocation into the nucleus.
As a result, WDR74-induced malignant transformation of lung cancer cells corresponds with β-catenin re-localization. Following that, the accumulation of β-catenin in the nucleus triggered the expression of downstream Wnt-responsive genes, resulting in uncontrollable cellular activity. Wnt-responsive genes such as c-myc and cyclin D1 have been linked to cell proliferation, whereas caspase 3, caspase 9, and MDR1 have been linked to chemo-resistance and death. WDR74 had regulatory impacts on these genes. In lung cancer cells, WDR74 influenced several biological processes in lung cancer cells by modulating these genes. In brief, WDR74 promotes lung cancer development and metastasis by increasing nuclear β-catenin accumulation and activating Wnt-responsive genes [11]. The objective of our research was to compare lung cancer protein sequence i.e. WDR74 protein homo sapiens sequence with the reference sequences; analyzing the local and global similarity between sequences by comparing protein sequence to sequence database, multiple sequence alignment and construction of phylogenetic tree.
II. MATERIAL AND METHODS
In this study, National Center for Biotechnology Information (NCBI) was used to retrieve the sequences. NCBI holds a number of databases related to biotechnology and biomedicine, and is a valuable asset for bioinformatics tools and services. GenBank, a database for DNA sequences, and PubMed etc. are some facilities provided by NCBI. After that, BLAST (Basic Local Alignment Search Tool) software was used for determining sequence similarity. BLAST only does "local" alignments. There is several BLAST variants available used in various sequence comparisons however in this study BLASTP and SMART BLAST was used. SMART BLAST analyses the protein query and generates a brief summary of the top five protein matches from well-studied reference species in the landmark database and it cannot discover five matches in the landmark database, it will utilize the protein non-redundant (nr) database. Conserved Domain Database matches to the query were also seen by SMART BLAST. Apart from it, software used for multiple sequence alignments were Clustal Omega and COBALT. Multiple sequence alignments are required in the majority of bioinformatics investigations that compare homologous sequences. Clustal Omega was primarily intended to align protein sequences; it did take nucleotide sequences as input. COBALT is a constraint-based alignment tool for multiple protein sequences with a broad architecture that use progressive multiple alignments to incorporate pairwise constraints from many sources into multiple alignments. It is part of the NCBI C Toolkit. It does not try to employ all accessible constraints, but rather a high-scoring consistent subgroup that can vary as the alignment progresses [12]. Following this, ORF Finder was used to scans the sequence for open reading frames (ORFs). Through this ORF range as well as the protein translation for each ORF was found.
III. RESULTS AND DISCUSSION
After retrieving the WDR74 protein sequence from NCBI, it is transformed into FASTA format and used as a query sequence in BLASTP. BLASTP compares a protein sequence with a database of protein sequences [13]. The BLASTP result displayed a list of hundreds of sequences in the description that were similar to the query sequence. The E value in the study of the WDR74 protein (WD repeat-containing protein) was zero, indicating that the query and subject sequences were better aligned.
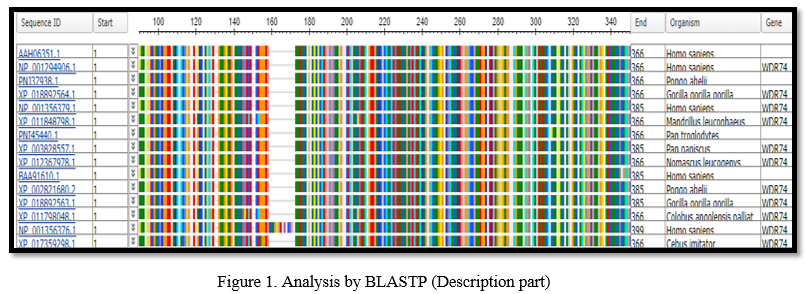
Query coverage was 100%, which indicated that the lengths of the query and subject sequences were the same. According to Per. Ident, the query sequence and WD repeat-containing protein 74 isoform 2 [Homo-sapiens] have a 100 percent similarity, whilst other sequences have a similarity ranging from 94.29 percent to 98.91 percent.
From the graphical summary and the description list it was observed that the hundred sequences have significantly more important alignments (with E-values of 0.0). The graphical summary showed alignments as colored boxes of database matches to the Query sequence and it showed red color indicating the greatest alignment scores (greater than or equal to 200), except NOP seven associated protein 1 [Macaca Fascicularis] that showed 93% similarity and the remaining sequence 7% showed gaps and mismatches to the actual sequence. After that, Structural tetrad on conserved domain WD40 superfamily was observed. 32 of 40 of the residues that compose this conserved feature have been mapped to the WDR74 sequence. Following this, three different sequence of same length were retrieved from NCBI, and was used in CLUSTAL OMEGA to observe the similarity among them. Clustal Omega used an alignment engine to align profile hidden Markov models (HMMs). This considerably improves Clustal Omega's accuracy [14]. It was found that certain sequences contain the same nucleotide and are conserved, as indicated by (*), but many sequences have several gaps, as indicated by (-). More gaps in the sequences suggested a higher likelihood of mutation. Following that, the constraint-based multiple sequence alignment method was applied. The query sequence was retrieved from NCBI in FASTA format and multiple aligned using COBALT. It does not try to employ all accessible constraints, but rather a high-scoring consistent subgroup that can vary as the alignment progresses [12]. The COBALT result emphasized the amino acid residues among the sequences, giving us insight into which aino acid is present in the sequences that are common.
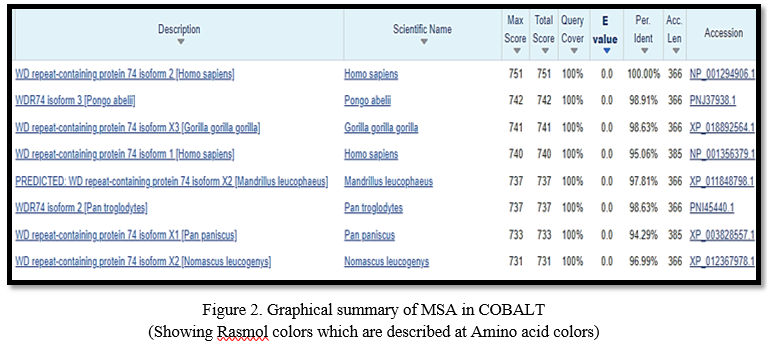
COBALT's graphical representation emphasized sequences that matched the query sequence by showing amino acids in different colors and providing additional information such as the organism's name to which the sequence belongs, their sequence ID, and the gene it possesses. The WDR74 query sequence showed similarity with sequences from Homo sapiens, Gorilla, Mandrillus leucophaeus, Pongo abelli etc. Following that, a phylogenetic tree was constructed using SMART BLAST
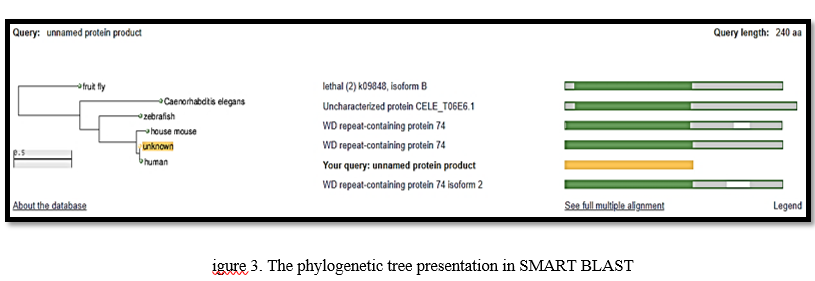

In this phylogenetic tree, the query is colored yellow and the matching sequences are from the reference database and were marked green. Deletions in the multiple sequence alignment appear as white gaps, and regions that did not align with the query sequences were marked as grey. The query sequence showed similarity in sequence with Caenorhabditis elegans (Mitogen-activated protein kinase mpk-1), fruit fly (rolled, isoform C), zebrafish (mitogen-activated protein kinase 1), Mus musculus (mitogen-activated protein kinase 1) and humans (mitogen-activated protein kinase 1). We found the frame containing mutations using ORF finder.

It showed six horizontal bars, each one representing a different reading frame. In each orientation of the DNA, there were three possible reading frames. As an outcome, the DNA sequence revealed six possible reading frames in all (six horizontal bars). The potential reading frames on the opposite strand are +1, +2, +3, and -1, -2, and -3. As a result, the reading frame with the most alterations was bolded at the top by the software, i.e. ORF4. We then tracked the start as well as end codons in the ORF4 by visualizing its sequence.
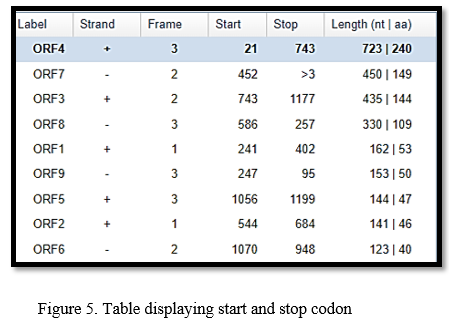
The result in the table showed that ORF4 has main mutations and it was observed that in ORF4, the start codons is at 21, 193, 286, 427, 718 position and stop codon is at 743 position
Conclusion
Our research findings support a possible causal link between WDR74 and lung cancer risk, and they imply that some genetic variables and some biological processes may play a role in the development of specific cancers. When compared to individual investigations, bioinformatics analysis can produce more trustworthy and accurate screening findings by overlapping pertinent data. Clinical studies and additional platforms\' gene chips are required to corroborate the findings of the discovered potential genes in lung cancer.
References
[1] R. J. Cersosimo, “Lung cancer: a review”, Am J Health Syst Pharm, Vol.59, pp. 611-42, Apr, 2002. [2] P. Jaggi, “A Review Article on Lung Cancer Diagnosis & Treatment”, JMAHS, Vol.6, pp. 1-9, 2017. [3] M. B. Schabath and M. L. Cote, “Cancer Progress and Priorities: Lung Cancer”, Cancer Epidemiol Biomarkers Prev., Vol. 28, pp. 1563-1579, Oct, 2019. [4] H. Lemjabbar-Alaoui, O. U. Hassan, Y.W. Yang and P. Buchanan, “Lung cancer: Biology and treatment options”, Biochim Biophys Acta., Vol.1856, pp. 189-210, Dec, 2015. [5] P. Rubin and J.T. Hansen, “Tnm staging atlas with oncoanatomy”, Lippincott Williams and Wilkins, 2012. [6] M. P. Curado, B. Edwards, H. R. Shin, H. Storm, J. Ferlay, M. Heanue and P. Boyle, “Cancer incidence in five continents” Vol. IX, pp.896, 2007. [7] A. J. Alberg, M. V. Brock, J. G. Ford, J. M. Samet and S. D. Spivack, “Epidemiology of lung cancer: Diagnosis and management of lung cancer” 3rd ed., American College of Chest Physicians evidence-based clinical practice guidelines, Chest, Vol. 143, pp. e1S-e29S, May, 2013. [8] A. El-Telbany and P. C. Ma, “Cancer genes in lung cancer: racial disparities: are there any?”, Genes Cancer, vol. 3, pp. 467-80, Jul, 2012. [9] J. Liu, M. Zhao, B. Yuan, S. Gu, M. Zheng, J. Zou, J. Jin, T. Liu and X. H. Feng, “WDR74 functions as a novel coactivator in TGF-? signaling”, Journal of Genetics and Genomics, vol. 45, pp. 639-650, 2018. [10] Y. H. Lo, E. M. Romes, M. C. Pillon, M. Sobhany and R. E. Stanley, “Structural Analysis Reveals Features of Ribosome Assembly Factor Nsa1/WDR74 Important for Localization and Interaction with Rix7/NVL2”, Structure, vol. 25, pp. 762-772, May, 2017. [11] Y. Li, F. Chen, W. Shen, B. Li, R. Xiang, L. Qu, C. Zhang, G. Li, H. Xie, V. L. Katanaev and L. Jia, “WDR74 induces nuclear ?-catenin accumulation and activates Wnt-responsive genes to promote lung cancer growth and metastasis” Cancer Lett., vol.471, pp. 103-115, Feb, 2020. [12] J. S. Papadopoulos and R. Agarwala, “COBALT: constraint-based alignment tool for multiple protein sequences. Bioinformatics”, vol.23, pp. 1073-9, May, 2007. [13] G. Syngai, P. Barman, R. Bharali and S. Dey, “BLAST: An introductory tool for students to Bioinformatics Applications”, Kenean Journal of Science, vol. 2, pp. 67-76, Dec, 2013. [14] F. Sievers and D. G. Higgins, “Clustal Omega for making accurate alignments of many protein sequences”, Protein Sci., vol.27, pp. 135-145, Jan, 2018.
Copyright
Copyright © 2022 Navjot Kaur Virk, Uma Kumari. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET43175
Publish Date : 2022-05-23
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

