Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- Conclusion
- References
- Copyright
A Review of Left Ventricle Segmentation Techniques
Authors: Anirudh P Kalghatkar, Darshan Sudheer Amadalli, Uday D, K Shashank, Dr. Kavitha K S
DOI Link: https://doi.org/10.22214/ijraset.2023.49169
Certificate: View Certificate
Abstract
The creation of automated techniques for the identification and segmentation the left ventricle (LV) in medical pictures has received more attention in recent years. This is partly caused by the rising incidence of cardiovascular disease and the demand for more precise and effective LV evaluation techniques. The considerable variation in LV form and size, as well as the presence of other structures like the right ventricle (RV) and left atrium in the same pictures, provide a number of obstacles for LV recognition and segmentation. In addition, other surrounding structures, such as the ribs, frequently block the view of the LV myocardium, the muscular wall of the heart. Despite these difficulties, several automated techniques have been put out lately, with varied degrees of effectiveness. We evaluate the current status of automated LV detection and segmentation in this research and provide an improved method for LV detection and segmentation. We test our strategy using a sizable datasets of cardiac MRI images, and we contrast the outcomes with those of other approaches.
Introduction
I.INTRODUCTION
According to the World Health Organization, cardiovascular diseases (CVDs) are the most lethal condition that endangers human life [1]. According to the American Heart Association, if cardiovascular illnesses can be averted by efficient early detection, human life expectancy can be increased by 10 years. Early CVD identification is therefore essential for choosing the right treatment to prolong human life and lower the death rate.
Cardiac MR imaging uses computerised technology that combines a strong magnetic field and radio wave to create detailed pictures of the anatomy of the heart. To diagnose underlying CVDs, cardiac MR imaging must identify various parameters of the left ventricle (LV) and right ventricle (RV). Therefore, it is essential to segment the LV and RV using MR imaging (MRI) before beginning the quantification process. Multiple research have created computerised systems using that can segment LV and RV since manual segmentation of MRI is a demanding process that can lead to human mistakes. The exterior and inner cardiac structures, as well as the difference in RV and LV size and shape between patients, are problems that these suggested methods must deal with. There has been significant advances in the Left Ventricle Segmentation methods And some methods are performed on MRI as well as ECG images of the heart to measure on the performance of these methods. We have prepared this survey paper based on mainly the dice score. In order to ensure that there is certain uniformity with respect to measuring the accuracy of the model, We have used Dice Metrics to ensure that all the proposed methods are studied carefully.
II. LITERATURE SURVEY
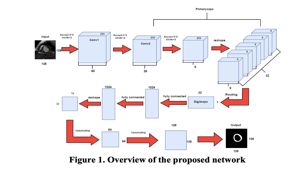
Jiaxiu Dong [et al] [1] discusses an approach to extract LV myocardial borders[Fig 1]. The proposed neural network consists of two conv layer, two capsule layer, one fully connected layer and two-deconv layers. The conv layer is the first layer that is used for feature extraction from the input image, It utilises a ReLU activation function. The First capsule layer is used to extract the low level image information that exists during this operation. A total of 5,308,416 parameters exists for this layer. While the Digital Capsule Layer outputs a vector for which, using the dynamic routing method we find the segmentation of the input image. The fully Connected layer is used to ensure that the features extract from the target region are as accurate as possible since the distances between pixels has undergone changes. In addition to this the Fully Connected Layer utilises a ReLU activation function. Finally in the first deconvolutional layer, The output from the second fully connected layer is reshaped and is taken as the input for the first deconv layer whose output are activated by the ReLU function further the first deconvolutional layer’s output is then fed into the last deconvolutional layer whose output if the resulting segmented image. The kernel of these two deconvolutional layer is said to be of 10x10. The average dice coefficient of this method is said to be 0.9417.
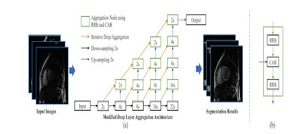
Zhongyu Li et al [2] discusses about a Deep Layer Aggregation method with improvements in the up-sampling path . A DLA-34 is chosen as the backbone for the networks. DLA uses two architectures namely Iterative Deep Aggregation(IDA) and Archical Deep Aggregation (HDA). Aggregation starts at the shallowest, smallest scale in iterative deep aggregation (IDA), and then iteratively merges deeper, larger stages. To learn richer combinations that cover a larger portion of the feature hierarchy, shallower and deeper layers are coupled using HDA. The DLA uses IDA as a form of skip connection that progressively aggregates for this model. The proposed improvement in the upsampling path involves the use of (i)Channel Attention Block (CAB) that varies the weight of features in each of the layers to improve consistency. (ii) Refinement Residual Block(RRB) that helps in combining the features of information that is being received from different input channels. The proposed method performed with a dice score of 94.2(±5.5) for the Left Ventricle.
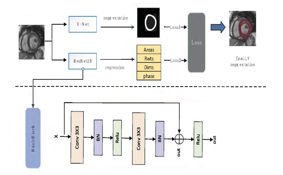
Zhi Liu et al [3] proposes a Multi Constrained Segmentation method based on multi task learning to perform left ventricle segmentation. In this method, Initially a U-net model is used first to segment the left ventricle to establish a standard for the MC-Seg to improve upon. The actual proposed method adds a ResNet 18 network to estimate various cardiac parameters including regional wall thickness, myocardial areas etc. When the multi task constraints are added to the network, It forces the network to learn these features to establish cardiac boundaries in between epicardium and endocardium. It was also observed that the repeated training of the multi task learning network helped reduce overfitting and improved the overall ability to segment the given mri image. A dice score of 0.886 was obtained from this proposed method, which was a significant improvement from the base U-net’s dice score of 0.845. Given below is the architecture of the proposed model :
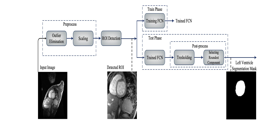
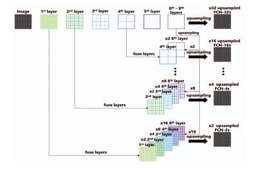
Mina Nasr-Esfahani et al [4] discusses a Fully Convolution Network method. The dataset obtained first has to be cleaned if any outliers present in the form of pixel intensity are removed by substituting it with a value absent from the set of outlier’s values. The region of interest extraction process consists of multiple stages wherein the first stage measures the difference in intensities of pixel between two successive frames followed by averaging the absolute difference causing the heart to be more salient. The resulting salient map based on a certain threshold is obtained within a square box which is now determined as ROI. A Fully Convolutional Neural Network is used which consists of layers same as a CNN except for a deconvolution layer in the last layer. However in this paper a set of three different types of FCNs are used, FCN-32s, FCN-16s, FCN-8s which differ by deconvolution feature maps. The output consists of a probability map on which a certain threshold is set using Otsu Algorithm[5] and in some cases may need further post-processing. One with the maximum roundness property is chosen as the Left Ventricle. Of the total of 5011 images, A 1000 is considered as test sets while the rest are training sets. When compared with existing methods, The current method has attained a dice score of 87.24% . However there still needs to be further work done to enhance the existing CNN architecture to obtain improved accuracy. Given below is the block diagram of the architecture:
Roshan Reddy Upendra et al [5]has explored the possibility of using GANs framework along with U-Net and comparing their result with that of a standalone U-Net. One such GAN is the SegAN consisting of two networks mainly: segmentor based on FCN that shows a probability map and is responsible for maximizing the loss function. Another network is that of a critic which requires two inputs - the image with actual labels and the other input consisting of image with predicted labels from the first network and this network deals with minimizing the loss function. Here three different sets of U-Net Architecture are used. Firstly the original U-Net[24], Second The one where the encoder-decoders are used as segmentors(U-Net A)[25] and Finally a redesigned UNet(U-Net B)[26]. The training of these segmentor and critic networks is based on back propagation and using cross entropy as a loss function. The results of the three networks are compared when used as a standalone U-net with the resulting standalone U-Net ‘s output as the segmentor and the downsampling section of the U-net as a critic. It was found that SegAN + U-net B shows promising results with a dice score of 95.87%.
Wei-YEN et al [6] proposed a faster R-CNN method to extract and recognize the aspects of the left ventricle, The tasks of segmenting and tracking the left ventricle is looked after by the Active Shape Model. To help deal with noises in the images, an enhanced anisotropic diffusion filter is introduced which utilizes 8 neighborhoods with varied weightings including α and β params. In the end a smoothing and a sharpening procedure are incorporated as well.
A pre-trained CNN network based on ZFNet is selected, Similarly a pre-trained ZFNet model is used to initialize the RPN for the faster R-CNN. Once the left ventricle has been identified from the Active Shape Model is used to segment and track the left ventricle. The Improved ASM model from [27] is used. The matching functions used Fig[i] are calculated by iterating and adjusting the shape parameters. In this study using faster R-CNN the task of finding the initial search position for the ASM algorithm is eliminated as it is located automatically from the R-CNN. A total of 30 series of ultrasound images are used each consisting of at least 1500-5000 images with the size of 636x434 were used in training the model. The dice score obtained with a training set of 600 images was obtained around 91.88%.
From YIHUA LAN et al [7] we find that a deep learning model was employed for a rough LVS on cardiac image. Further a double snake model was utilized. In the end, The optimal epicardial boundary obtained by radial region growing method. The dataset was obtained from MICCAI. A SegNet based approach was used for the process of segmentation of the short-axis images, The results from this model were filtered for myocardial contours. To further delineate the resulting image into its endo and epi-cardial boundaries following snake model was proposed
Lt+1 = Lt + γ(αE ∗ FE + αB − αC ∗ FC)
The resulting endo and epi cardial contours found by the snake model along the epicardial contour region acquired by the previous slice images were used to create a binary mask that would be used to calculate the epicardial contour. Once the ground truths were established, The quality of the endo- and epi- cardial contour was determined based on their mean distance from the ground truth, For a good contour the maximum threshold was less than 5mm. The resulting Dice metric from the Endo and Epi- Cardial were found to be 0.96 and 0.97 respectively. Though this method was found to be reliable, There still remains certain challenges when it comes to outliers that may exists for which further study is needed. Schematic diagram of the process is given below :
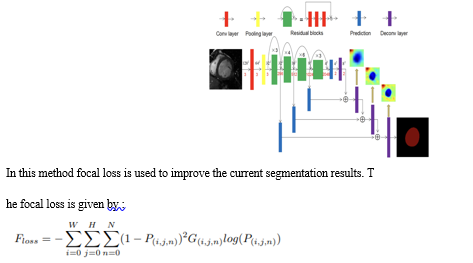
Mingqiang Chen [et al] [8] proposes a a focal residual network (FR-Net) that combines skip connections and focal loss that is commonly used in object segmentation. A ResNet is utilised as a backbone of this network. Whose architecture is shown as follows:
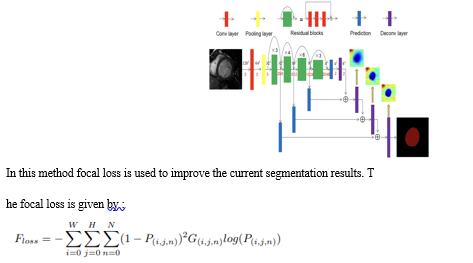
An alternative training strategy is also put forward where we would first optimize the parameters of this network using focal cross entropy loss functions to converge, Then we switch to the dice loss function parameter to optimize the function. This process would keep on continuing till the losses no longer change. A dice metric score of 0.93 was obtained in the FR-Net using the Sunnybrook public dataset.
Ayush Goyal et al [9] discusses that MRI images can be segmented using part based growing in ITK SNAP. Then the cardiac ventricles are taken out from the 3D cardiac MRI image stack of the entire heart beat cycle. Different constraints such as myocardial thickness can be found out which can be further analyzed by physicians for safety of the patients. Image processing has been done through the initialization of seedpoint and developing of locale .Computational reconstruction has been done in such a way that the whole MRI picture dataset is constructed into VTK design in ITK-SNAP software format which is used for division .The 3 dimensional clinical data can be visualized. Region of interest can be found out along with active contour can be used for segmentation strategy is fast and it is free of iterations which are expensive in terms of computation and requires initial seed point.
The method suggested by Xianbin Zhu[et al][10] a pixel based FCM method with level set method is utilized. The FCM divided the MRI images into two different clusters of foreground and background. The MRI images are labeled based on the Connected Component Label method which is further processed by the proposed convexity level set method. For clustering the MRI images, each pixel is split into multiple clusters and their membership levels decides which clusters the pixels belong to.To further locate the left ventricle region obtained from the FCM method a label selection process is performed, This consists of two parts where firstly, multiple connected components are marked followed by a process to filter out noisy labels. A Convexity preserving level set method is used to enhance the segmentation of left ventricle.The gradient equation is as follows:
This approach has been tested on datasets of MICCAI 2009 then algorithm has been applied to 300 images .The CPLS algorithm suits well because dice score was 0.90 takes 1.02 sec with less number of iterations
R.Sreemathy et al[11] proposes that during MRI of the heart will be divided from apex to base into different levels 20 frames will be considered per level left ventricle is extracted from each and every level to improve the accuracy of segmented boundaries of left ventricle a anisotropic filtering along with gaussian smoothing has been used 2 frame from every level having less and more area has been separated , lesser area frame can be noted as end of systole phase while that of larger area frame can be noted as end of diastole phase. Combining the area of both the net area of left ventricle can be calculated. The result were analyzed with regression technique and bland altman plot The regression equation is given by Y=0.1527+0.9993X The Bland Altman plot has been done using snake algorithm
S.Molaei et al[12] made a comparison between manual segmentation and automated segmentation In this paper Deep Convolutional Neural Network (DCNN) is utilised for LV Segmentation. This algorithm produces a binary label image that assigns Left Ventricle wall pixels a score of 1 and other pixels a tag of 0. MatConvNet is chosen as the backbone for the neural network. The structure for imagenet has 31 layers followed by alternative convolution,RELU,pooling.The last 40th layer gives the segmentation prediction image. Using the standard gradient descent backpropagation algorithm, the network is trained with the softmax log-loss function. To aid the DCNN in unbalanced pixel distribution for two steps has been approached firstly, extracting the Region of Interest (ROI). Secondly, weighting the image dependent on the relative class frequencies. Followed by cost sensitive pixel classification in which number of pixel which come under left ventricle will be smaller A matrix of weights has been applied to the input image.Followed by filter initialization has taken place between random initialization and Gabor initialization then these 2 methods were compared with DCNN structure.Testing has been done with 150 MRI images from the dataset. The performance metrics has been used to compare the result between gabor filter and random filter gabor filter gives the highest performance metric value in terms of accuracy which records 0.973 Specificity 0.984 sensitivity 0.841 mean accuracy 0.902
O.Grebe et al[13] states that endocardial segmentation has been done with the help of the two phases Segmentation phase and Refinement phase. In the first phase Bottom up multi scale analysis has been used by utilizing morphological scale space. With response the gray level structures which are next to the endocardial border are discovered and the approximate dividing lines are secured followed by a refinement phase that asserts the previous knowledge about local structure around defined points within the shape boundary. The proposed method was able to segment the inner structures in 91% tested cases from 220 MR images
Vinayak Ray et al[14] has explored that in MRI images has been undergone through the series of the algorithms such as Adaptive k-means clustering to determine the number of clusters, Morphological hole fill, Local Otsu thresholding, Connected component labeling,Noise removal, Heuristic segmentation, Boundary detection. The resulting clusters are then passed on to the morphological hole filing to check no holes are present in the cavity this image is passed on to the Local Otsu thresholding to separate foreground from background clusters then this is followed by next layer Connected component labelling based on the connectivity threshold image is grouped into region of pixels and then noises are removed And then it is followed by Heuristic segmentation, Boundary detection in which the boundary points of the left ventricle can be determined. Automatic segmentation records correlation of 0.992 for frame 1 and 0.993 for frame 3 This approach is quick as it is free of human interventions
A CNN based left ventricle segmentation using 2D ultrasound images was proposed by Erik Smistad et al[15],based on CNNs are being researched for the segmentation of LV ultrasound images. The outpatient clinic received 1,500 ultrasound recordings from 100 individuals. These recordings included around one hundred thousand 2D picture frames. The left ventricle is represented as a cubic hermite spline in this segmentation approach. A Kalman filter is used to estimate and forecast all of these transition state parameters throughout time (KF).By using pre-trained CNN with a previously described automated Kalman filter (KF)-based segmentation approach, the requirement for manual annotation can be eliminated. The results indicate that by merely training using created data, a CNN may attain equivalent accuracy to the automated technique. Stochastic gradient descent with ten epochs, learning rate one percent and momentum 0.9 was employed for network training.The dice similarity coefficient was 0.86 ,0.06 for the CNN vs 0.87 ,0.06 for the KF technique, while the Hausdorff distance was 5.9 ,2.9 mm for the CNN versus 7.5, 5.6 mm for the KF method.
Vasily Zyuzin et al[16] discussed on U - net architectures are compared for segmenting the left ventricle endocardial boundary on two-dimensional ultrasound images. The network architectures considered are U-NET, WIDE U-NET AND UNET++.The data collection includes video sequences of full cardiac cycles from ninety four people from three groups: those without pathologies, those with ischemic heart disease, and those with dilated cardiomyopathy. The issue of automated segmentation and tracing the Left Ventricle on EchoCG pictures is a real-world challenge. They recommended employing multilayer CNNs for pixel categorization in images to solve the problem. Every fold can be used as a validation subset. The remaining components are used to train a model. As a result, they created five models for each architecture and tested them on a validation subset. To increase segmentation quality, new models based on artificially augmented data have been developed. The main difference between Unet++ and Unet is enhanced skip connections in the Unet architecture via convolution blocks. Skip connection enables us to transport information from encoder to decoder while preserving minor features of the ultrasound picture. It is essential to accurately recognize internal borders. Net++ was examined in contrast to Unet and Wide Unet. The evaluation matrix for the LV segmentation is done using Dice Coefficient which was found to be 91.20% in this scenario. Unet++ surpasses the Unet and wide Unet designs for segmentation of LV.
Manuel Pérez-Pelegríet al[17]. presented a modern convolutional neural network design that integrates U-net and PSP modules (PSPU-net). Their dataset included three hundred and ninety nine short-axis MRI images. All of the photos were randomly divided into three sets: training (260 cases,),for testing (40 cases,), and testing (99 cases).Two distinct U-net designs were created and trained. The primary design is 3D Segmentation using a pre-trained U-net on the collected data and the other is PSPU-net. There are some major modifications in the PSPU-net. Despite inputs similar to the 3D U-net (176x176x3), the convolutions in this design are 3x3x1,calling it a two-dimensional U-net because it only utilizes 2D information. The loss function training record for both networks functioned similarly, with identical training and validation loss evolutions. Another significant finding is that the PSPU-net training varied less in terms of its loss function making it more stable. Overall, the proposed method has a higher dice scores and lower standard deviations, making it more resilient. In addition, because the PSPU-net is a smaller network (approximately one-third of that of U-net ),hence is more efficient. For a test set of 99 cases, the proposed method obtained a dice score of 0.910.
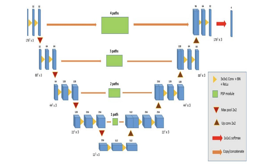
Hisham Abdeltawab et al [18]. have presented a modern framework for automatic segmentation of the epicardial and endocardial walls of LV using a fully convoluted Unet neural network directly from cardiac images. They were able to ease the class imbalance presenting a single loss function that accounts for the data fluctuation. The new feature reduces the impacts of binary cross-entropy by enhancing the final segmentation’s sensitivity and specificity.. Development of an efficient and accurate U-net-based deep learning model to automatically determine the epicardial and endocardial walls of all LV slices directly from CMR images. Therefore, this technique requires no pre-processing to detect LV. Proposed LV segmentation structure. Part (a) contains CMR photos used as input. Part (b) shows the U-net architecture
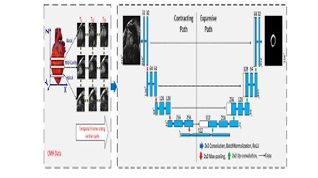
Using 30 distinct short-axis CMR datasets, the suggested pipeline was evaluated and appraised. Each set has 7-15 slices/sequences covering the patient’s left ventricle (apical, medial, and basal). The training set includes 3700 pictures from 16 movie. Compared to the standard loss function, our technique showed promising accuracy for epicardial and endocardial wall segmentation (Dice 0.94 or 0.96)
Vinayak Ray et al [19], proposes a method to accomplish accurate and quick segmentation of the left ventricle, The technique combines Image-Based Fuzzy C-Means Clustering with Connected Component Labeling. In Image Based Fuzzy C-Means Clustering, the approach employed in this work, considers both intensity values and spatial information to increase segmentation accuracy. The technique of Linked Component Labelling is used to detect and label connected areas in a picture.
In this case, the left ventricle is identified using this method from the MRI. By combining these two techniques, the method is able to accurately perform the task of segmentation. The results show that the proposed method achieves a high level of accuracy in segmenting the left ventricle, with a Dice similarity coefficient of 0.92.
Xinyi Li et al [20],presents a novel approach for automatic segmentation of the LV using a multi-scope CNN. The method used based on the idea of making use of multiple CNNs with different receptive fields to capture features at various scales in the images, which is known as multi-scope CNN. The proposed approach consists of 2 stages: The first stage involves the use of multi-scope CNN. The second stage involves use of connected component analysis to the output of the first stage to extract the left ventricle region. The multi-scope CNN is trained on a dataset of cardiac MRI images and it is composed of multiple branches, each one with a different receptive field, which allows it to capture features at different scales. These branches are combined together to generate the final probability map of the left ventricle. The method’s performance was examined using a dataset of cardiac MRI images, and the findings suggest that it can achieve high accuracy in segmenting the left ventricle, with a Dice coefficient of 0.92. The proposed method has several advantages over other traditional methods for left ventricle segmentation. It is fully automatic and does not require any manual intervention, which can save time and decrease human error. Furthermore, it may separate the left ventricle from the overall cardiac cycle, providing more accurate and full information on the heart’s activity.
Zewen Li et al [21]. discusses an approach of combining CNN with a model of active contours and tensor voting. The method is designed to overcome the limitations of traditional methods that rely solely on CNNs or active contour models, and achieves high accuracy in segmenting the left ventricle.The method consists of three stages: Firstly, the CNN is used to generate a probability map of the left ventricle, in the second stage, an active contour model is added to the probability map and in the third stage, Tensor voting is utilized to boost segmentation accuracy even more. The active contour model is designed to take into account the shape and topology of the left ventricle to improve accuracy. A Skip architecture used in this method.
The tensor voting uses a series of voting fields to determine if the left ventricle exists or not in an MRI image. This can help to overcome any errors or inaccuracies that may have been introduced by the CNN or active contour model.The proposed method’s performance was tested and a Dice similarity coefficient of 0.94 was obtained.
Li Kuo Tan et al[22], proposes a method that usually circular LV endocardium can be easily parameterized into a polar radial system. The LV border was then segmented using neural network regression of the radial distances rather than the traditional categorization of each pixel. This solution decreases the problem’s dimensionality, allowing for a smaller network design while achieving decent results without the need for costly post-processing. They created two convolutional neural networks (CNN), one for LV localisation and one for assessing endocardial radius. The networks were trained using hundred datasets from the MICCAI 2011 competition and evaluated using forty nine datasets from the MICCAI 2009 challenge. The networks obtained an average Dice metric of 0.88, a perpendicular distance of 2.30 mm, and 97.9% good contours.
Anupama Bhan et al[23] has used fuzzy C-means, a pixel-based classification approach and linked component labeling to segment the left ventricle. Each frame’s average computation time is 0.7 seconds because of the complementarity between the two methods. Employing a defined relevance factor , the fuzzy c-means (FCM) method divides the input cardiac image’s in pixels into a collection of c fuzzy clusters.
Every partition matrix component, which is membership level wqc, in the method’s definition of cluster centres determines the proportion of pixel xi that is a part of cj. As a consequence, compared to iteration-based methods, this technique’s left ventricle segmentation computation time is significantly shorter. Based on correlation coefficient, When compared to manual segmentation, the accuracy of the automatic segmentation method was evaluated. The automated and manually traced LV borders have a correlation coefficient of 0.932, which is regarded as clinically significant. Given that the variation between the inside and outside walls and the solidity of the blood pool can be found out, another criteria for clinical diagnosis, the future experimentation can be expanded to segment the left ventricle’s outside wall, also known as the epicardium. Fuzzy clustering is used to compare automatic segmentation with manual segmentation, with a correlation between automatic ASI and manual MS2 of p (ASI, MS2) = 0.932.
III. DICE COEFFICIENT
The Dice coefficient can be used to compare the pixelwise agreement between a predicted segmentation and its corresponding ground truth.
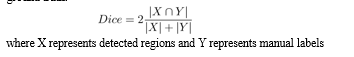
Conclusion
This literature survey contains different methods and approaches used to precisely segment the left ventricle of the heart. A total of 30 papers were considered based on the proposed methods used for detecting segmenting the left ventricle from the MRI images and their dice scores were compared to find out the best performing method. This paper aims to highlight the different methodologies that have been used to improve the accuracy with the left ventricle segmentation. Publicly available datasets such as Sunny Brooks MRI, MICCAI were used. Various techniques were utilized for the Data Pre-processing steps as well during Data Augmentation. Finding the Region of Interest was the most important process before training the model as it is necessary to ensure that only useful information is being fed into the model. We found that two proposed methods had desirable results based on their dice score. Firstly, Hisham Abdeltawab[et al] ’s proposal resulted in a dice score of 0.96 for endocardium and 0.97 for epicardium. Secondly, YIHUA LAN[et al] yielded similar results for both endo epicardium. Their performance were more satisfactory compared to the methods mentioned in the survey.
References
[1] Jiaxiu Dong, Chang Liu, Cong Yang, Nan Lin, and Yangjie Cao. 2018. Robust Segmentation of the Left Ventricle from Cardiac MRI via Capsule Neural Network. In Proceedings of the 2nd International Symposium on Image Computing and Digital Medicine (ISICDM 2018). Association for Computing Machinery, New York, NY, USA, 88–91. https://doi.org/10.1145/3285996.3286016 [2] H. Abdeltawab et al., ”Automatic Segmentation and Functional Assessment of the Left Ventricle using U-net Fully Convolutional Network,” 2019 IEEE International Conference on Imaging Systems and Techniques (IST), Abu Dhabi, United Arab Emirates, 2019, pp. 1-6, doi: 10.1109/IST48021.2019.9010123. [3] Z. Liu, P. Li, J. Li, Q. Xie and X. Wang, ”Left ventricular full segmentation from cardiac Magnetic Resonance Imaging via multi-task learning,” 2021 IEEE 2nd International Conference on Big Data, Artificial Intelligence and Internet of Things Engineering (ICBAIE), Nanchang, China, 2021, pp. 71-75, doi: 10.1109/ICBAIE52039.2021.9390044. [4] M. Nasr-Esfahani et al., ”Left Ventricle Segmentation in Cardiac MR Images Using Fully Convolutional Network,” 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Honolulu, HI, USA, 2018, pp. 1275-1278, doi: 10.1109/EMBC.2018.8512536. [5] Upendra, R.R., Dangi, S., Linte, C.A. (2019). An Adversarial Network Architecture Using 2D U-Net Models for Segmentation of Left Ventricle from Cine Cardiac MRI. In: Coudi`ere, Y., Ozenne, V., Vigmond, E., Zemzemi, N. (eds) Functional Imaging and Modeling of the Heart. FIMH 2019. Lecture Notes in Computer Science(), vol 11504. Springer, Cham. https://doi.org/10.1007/978-3-030-21949-945 [6] W. -Y. Hsu, ”Automatic Left Ventricle Recognition, Segmentation and Tracking in Cardiac Ultrasound Image Sequences,” in IEEE Access, vol. 7, pp. 140524-140533, 2019, doi: 10.1109/ACCESS.2019.2920957. [7] W. -Y. Hsu, ”Automatic Left Ventricle Recognition, Segmentation and Tracking in Cardiac Ultrasound Image Sequences,” in IEEE Access, vol. 7, pp. 140524-140533, 2019, doi: 10.1109/ACCESS.2019.2920957. [8] M. Chen, L. Fang and H. Liu, ”FR-NET: Focal Loss Constrained Deep Residual Networks for Segmentation of Cardiac MRI,” 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), Venice, Italy, 2019, pp. 764-767, doi: 10.1109/ISBI.2019.8759556. [9] A. Goyal, D. Bathla, P. Sharma, M. Sahay and S. Sood, ”MRI image based patient specific computational model reconstruction of the left ventricle cavity and myocardium,” 2016 International Conference on Computing, Communication and Automation (ICCCA), Greater Noida, India, 2016, pp. 1065-1068, doi: 10.1109/CCAA.2016.7813900. [10] X. Zhu, M. Li and F. Shan, ”Segmentation of cardiac left ventricle from MRI images using a method based on FCM and convexity preserving level set,” 2019 International Conference on Medical Imaging Physics and Engineering (ICMIPE), Shenzhen, China, 2019, pp. 1-5, doi: 10.1109/ICMIPE47306.2019.9098235. [11] R. Sreemathy and R. S. Patil, ”Computer assisted analysis of left ventricle using level set and enhanced filtering in cardiac MRI,” 2012 38th Annual Northeast Bioengineering Conference (NEBEC), Philadelphia, PA, USA, 2012, pp. 349-350, doi: 10.1109/NEBC.2012.6207108 [12] S. Molaei, M. Shiri, K. Horan, D. Kahrobaei, B. Nallamothu and K. Najarian, ”Deep Convolutional Neural Networks for left ventricle segmentation,” 2017 39th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Jeju, Korea (South), 2017, pp. 668-671, doi: 10.1109/EMBC.2017.8036913. [13] H. El-Messiry, H. A. Kestler, O. Grebe and H. Neumann, ”Segmenting the endocardial border of the left ventricle in cardiac magnetic resonance images,” Computers in Cardiology, 2003, Thessaloniki, Greece, 2003, pp. 625-628, doi: 10.1109/CIC.2003.1291233 [14] A. Bhan, A. Goyal and V. Ray, ”Fast fully automatic multiframe segmentation of left ventricle in cardiac MRI images using local adaptive k-means clustering and connected component labeling,” 2015 2nd International Conference on Signal Processing and Integrated Networks (SPIN), Noida, India, 2015, pp. 114-119, doi: 10.1109/SPIN.2015.7095354. [15] E. Smistad, A. Østvik, B. O. Haugen and L. Lvstakken, ”2D left ventricle segmentation using deep learning,” 2017 IEEE International Ultrasonics Symposium (IUS), Washington, DC, USA, 2017, pp. 1-4, doi: 10.1109/ULTSYM.2017.8092573. [16] V. Zyuzin and T. Chumarnaya, ”Comparison of Unet architectures for segmentation of the left ventricle endocardial border on two-dimensional ultrasound images,” 2019 Ural Symposium on Biomedical Engineering, Radioelectronics and Information Technology (USBEREIT), Yekaterinburg, Russia, 2019, pp. 110-113, doi: 10.1109/USBEREIT.2019.8736616. [17] M. P´erez-Pelegr´? et al., ”PSPU-Net for Automatic Short Axis Cine MRI Segmentation of Left and Right Ventricles,” 2020 IEEE 20th International Conference on Bioinformatics and Bioengineering (BIBE), Cincinnati, OH, USA, 2020, pp. 1048-1053, doi: 10.1109/BIBE50027.2020.00177. [18] H. Abdeltawab et al., ”A Novel Deep Learning Approach for Left Ventricle Automatic Segmentation in Cardiac Cine MR,” 2019 Fifth International Conference on Advances in Biomedical Engineering (ICABME), Tripoli, Lebanon, 2019, pp. 1-4, doi: 10.1109/ICABME47164.2019.8940294 [19] V. Ray and A. Goyal, ”Image-based fuzzy c-means clustering and connected component labeling subsecond fast fully automatic complete cardiac cycle left ventricle segmentation in multi frame cardiac MRI images,” 2016 International Conference on Systems in Medicine and Biology (ICSMB), Kharagpur, India, 2016, pp. 36-40, doi: 10.1109/ICSMB.2016.7915082. [20] X. Li, Y. Wang, W. Yan, R. J. Van der Geest, Z. Li and Q. Tao, ”A Multi-Scope Convolutional Neural Network for Automatic Left Ventricle Segmentation from Magnetic Resonance Images: Deep-Learning at Multiple Scopes,” 2018 11th International Congress on Image and Signal Processing, BioMedical Engineering and Informatics (CISP-BMEI), Beijing, China, 2018, pp. 1-5, doi: 10.1109/CISPBMEI.2018.8633185. [21] .Z. Li, A. Lin, X. Yang and J. Wu, ”Left ventricle segmentation by combining convolution neural network with active contour model and tensor voting in short-axis MRI,” 2017 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Kansas City, MO, USA, 2017, pp. 736-739, doi: 10.1109/BIBM.2017.8217746. [22] L. K. Tan, Y. M. Liew, E. Lim and R. A. McLaughlin, ”Cardiac left ventricle segmentation using convolutional neural network regression,” 2016 IEEE EMBS Conference on Biomedical Engineering and Sciences (IECBES), Kuala Lumpur, Malaysia, 2016, pp. 490-493, doi: 10.1109/IECBES.2016.7843499. [23] A. Bhan, A. Goyal, M. K. Dutta, K. Riha and Y. Omran, ”Image-based pixel clustering and connected component labeling in left ventricle segmentation of cardiac MR images,” 2015 7th International Congress on Ultra Modern Telecommunications and Control Systems and Workshops (ICUMT), Brno, Czech Republic, 2015, pp. 339-342, doi: 10.1109/ICUMT.2015.7382454. [24] Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomed- ical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). [25] Xue, Y., Xu, T., Zhang, H., Long, L.R., Huang, X.: Segan: adversarial network with multi-scale l 1 loss for medical image segmentation. [26] Isensee, F., Jaeger, P.F., Full, P.M., Wolf, I., Engelhardt, S., Maier-Hein, K.H.: Automatic cardiac disease assessment on cine-MRI via time-series segmentation and domain specific features. In: Pop, M., Sermesant, M., Jodoin, P.-M., Lalande, A., Zhuang, X., Yang, G., Young, A., Bernard, O. (eds.) STACOM 2017. LNCS, vol. 10663, pp. 120–129. Springer, Cham (2018) [27] B. van Ginneken, A. F. Frangi, J. J. Staal, B. M. ter Haar Romeny, and M. A. Viergever, “Active shape model segmentation with optimal fea- tures,” IEEE Trans. Med. Imag., vol. 21, no. 8, pp.924–933, Aug. 2002.
Copyright
Copyright © 2023 Anirudh P Kalghatkar, Darshan Sudheer Amadalli, Uday D, K Shashank, Dr. Kavitha K S. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET49169
Publish Date : 2023-02-20
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

