Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- Conclusion
- References
- Copyright
Malaria Cell Detection Using Machine Learning
Authors: Mayank Singh, Rishabh Khurana, Parichay Jain, Ayush Verma
DOI Link: https://doi.org/10.22214/ijraset.2022.42831
Certificate: View Certificate
Abstract
Malaria is a major public health issue that affects people all around the world. The diagnosis of red blood cells contaminated with insects under a microscope by a skilled specialist is a typical way of diagnosing malaria. This method does not function effectively, and the diagnosis is made based on the test taker\'s experience and expertise. Malaria blood tests have been using automated imaging technologies based on machine learning for early detection. However, thus far, practical performance is insufficient. The convolutional neural network CNN is used in this paper to present an innovative and resilient machine learning model. for automatically distinguishing single cells in tiny blood smears from normal microscope slides, such as infected or non-infected with this virus. Our new CNN model\'s average accuracy for 16 layers is 97.37 percent based on ten-fold confirmation using 27,578 single-image pictures. In the same photos, the learning transfer model only gets 91.99 percent. All performance indicators, such as sensitivity (96.99 percent vs 89.00 percent), clarity (97.75 percent vs 94.98 percent), accuracy (97.73 percent vs 95.12 percent), F1 school (97.36 percent vs 90.24) percent, and Matthew\'s correlation coefficient, demonstrate that the CNN model outperforms the transfer learning model (94.75 percent vs 85.25 percent).
Introduction
I. INTRODUCTION
It is a serious human infectious illness. Malaria is caused by an insect protozoan called Plasmodium that may infiltrate human erythrocytes and produce a range of symptoms. Malaria infected 214 million individuals in 2014, killing over 438,000 people, according to the WHO. The annual economic effect is Malaria could have been prevented, controlled, and successfully cured if there had been a more accurate and effective diagnosis. The most common way to diagnose malaria is to detect abnormal blood smear in infected erythrocytes with trained microscopes. This procedure, however, does not perform well, and the quality of diagnosis is dependent on microscope expertise and understanding. Rapid
diagnostic techniques are also popular, even though they are more costly and give less information than microscopy.
In this exercise, we use an in-depth study method to detect red blood-borne red blood cells in a standard microscope slide prepared using standard methods. We use a convolutional neural network (CNN) model, which is an in-depth reading model designed for two-dimensional data reading such as photos and videos. Encourages the evaluation of basic physiological processes in the visual cortex of visual felines. The test encourages pattern recognition modeling to mimic visual cognitive processing The advantage of the CNN model is that its subsequent formation of learning layers. can be trained rigorously if the topology of the model is equal to the input element. The model has a lot of potential to use the local relationship of visual patterns (e.g., edges in a picture) to decrease the number of parameters that need to be read. This improves the accuracy of the feed forward-back propagation training process. Since in-depth learning can model very complex features, CNN provides a standard-purpose reading framework that does not require pre-configuration and fine tuning, which is more advantageous than regular class dividers.
Malaria is a life-threatening disease caused by Plasmodium viruses that attack the red blood cells (RBCs). Personally, detecting and counting parasitic cells in thick blood tests / thin film is still common, but it is a burdensome way to diagnose the disease. Its diagnostic accuracy has a negative impact on the internal / internal variability of the viewer, especially on large scale tests under resource-pressed settings. For image recognition applications, advanced computer-assisted diagnostic tools based on deep learning data, such as the convolutional neural network (CNN), have become the preferred structur. However, CNN suffers from high variability and may be exaggerated because of its sensitivity to training data variability.
Biology has several issues that have resulted from object discovery. Although there has been a lot of interest in models based on in-depth learning and their success in acquiring an object, the state-of-the-art models from competitions such as Image Net Large Scale Visual Recognition Challenge (ILSVRC) 1 and MS-COCO2 not yet widely used in image biological data. We're curious about finding something to identify cells and identify them stages of diseases such as malaria, in which self-examination of small ideas by trained specialists remains a gold standard. A solid solution would allow automatic cell division and calculation will also provide significant benefits due to faster and more accurate quantity results without human variability.
The detection of cell object in light microscope images presents special challenges. Like nature pictures, small pictures of malaria-infected blood the variation of light from the microscope, the shape of the cell, the density, and the color from the various sample preparations, and having the elements of an uncertain class (even artisans). However, unlike natural images, there is a lack of annotation data that is useful for training due to a lack of specialists, and class distribution is naturally severely imbalance due to the dominance of uninfected red blood cells (RBCs).
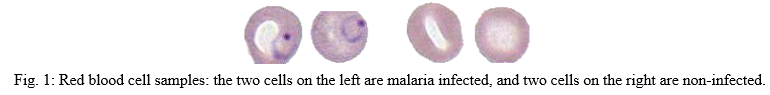
II. MOTIVATION FOR THE PROJECT
Malaria is a dangerous, contagious mosquito-borne disease caused by Plasmodium bacteria. The bite of female Anopheles mosquitoes transmits these insects. Although we will not go into detail about the disease, There are five different types of malaria. Now let's look at the importance of how deadly this disease can be in the next episode.
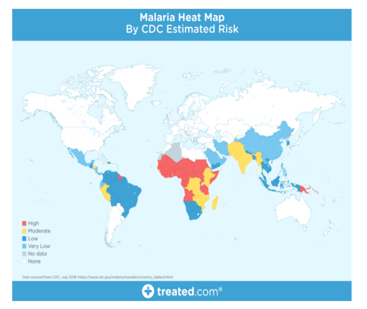
Clearly, malaria is rampant throughout the world, especially in tropical areas. The cause of this activity however depends on the nature and mortality of the disease. Initially when an infected mosquito bites, mosquito-borne parasites will invade your bloodstream and begin destroying the RBCs carrying oxygen (red blood cells). The first symptoms of malaria are often similar to the flu or the virus when you usually start getting sick a few days or weeks after being bitten by a mosquito. However, these deadly insects can live in your body for over a year without problems! Thus, delays in appropriate treatment can lead to complications and even death. So early and effective testing and detection of malaria can save lives.
The World Health Organization (WHO) has released a number of important facts about malaria that you can look at here. In short, malaria affects approximately half of the world's population, with more than 200 million cases reported and some 400,000 deaths from malaria each year. This gives us great inspiration to make malaria detection and diagnosis faster, easier and more effective.
Traditional methods of detecting malaria are time consuming, may produce inaccurate reports due to human error, and are difficult to diagnose. This encourages us to promote automatic detection of malaria using in-depth machine learning strategies and then using a web interface that leads to early detection which is quick, easy, and effective.
As we aim to develop an effective web-based detection for malaria cell, we look forward to coming up with a comprehensive CNN-based learning model that is expected to be simpler and more mathematically viable compared to most of the province. the art techniques discussed before that require a lot of training time. In particular, we make the following contributions: (a) designing and testing a CNN basic model with a standard or non-standard reading schedule and very little training parameters to distinguish viral and virus-free cellular images, automated in addition to the usual techniques used to improve model performance, and (b) the deployment of our most effective model in web application to facilitate easy and rapid malaria detection.
III. LITERATURE REVIEW
Machine learning and image analysis for detecting malaria in this research paper, a survey article on image analysis and machine learning methods to give an update on the latest development in automated malaria diagnosis with image analysis and machine learning has been written. Mahdieh Poostchia, Kamolrat Silamutb, Richard J. Maudebcd, Stefan Jaegera, George Thoma, “Image analysis and machine learning for detecting malaria”. Received 30 October 2017, Revised 7 December 2017, Accepted 19 December 2017, Available online 12 January 2018. [1]
Comparison of Detection Method on Malaria Cell Images In the image analysis process, one of the most important preprocessing steps is thresholding. This project will describe a few selected thresholding methods such as Wolf's method, Fuzzy C-Mean Algorithm's method, Bradley’s method, Bernsen's method, Triangle's Method and Deghost's Method. [2]
Clustering-Based Dual Deep Learning Architecture for Detecting Red Blood Cells in Malaria Diagnostic Smears In this research paper we learn about Dual deep learning architecture RBCNet, which combines U-Net with Faster R-CNN, provides a robust solution for detecting RBCs in blood smear images characterized by a small ratio of cell object size to image size.[3]
Automatic Detection of Malaria Parasites for Estimating Parasitemia S. S. Savkare Moze College of Engineering, University of Pune; S. P. Narote Sinhgad College of Engineering, University of Pune, Pune, India. In this research paper we find out, Malaria parasitemia is a measurement of the amount of Malaria parasites in the patient's blood and an indicator for the degree of infection.[4]
Applying Faster R-CNN for Object Detection on Malaria Images Jane Hung, Anne Carpenter; Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) Workshops, 2017, pp. 56-61. Deep learning-based models have had great achievement in object detection, but the state-of-the-art models have not yet been widely applied to biological image data. [5]
Automated Detection of Malaria Pigment in White Blood Cells for The Diagnosis of Malaria in Portugal A novel automated method (Cell-Dyn 3500) allows malaria diagnosis by detecting malaria pigment in white blood cells during routine full blood counts. In Portugal, 174 samples from 148 patients who presented to the emergency department were analyzed. Compared with microscopy the sensitivity was 95% and the specificity was 88%. [6]
Improving Malaria Parasite Detection from Red Blood Cell using Deep Convolutional Neural Networks. In this work, we conduct a series of experiments based on end-to-end deep learning to improve malaria classification from segmented red blood cell smears. Aimon Rahman, Hasib Zunair, M Sohel Rahman, Jesia Quader Yuki, Sabyasachi Biswas. [7]
Z. Liang et al., "CNN-based image analysis for malaria diagnosis," 2016 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 2016, pp. 493-496, doi: 10.1109/BIBM.2016.7822567.[8]
Deep Learning Based Automatic Malaria Parasite Detection from Blood Smear and Its Smartphone Based Application by K. M. FaizullahFuhad, Jannat Ferdousey Tuba, Md. Rabiul Ali Sarker, Sifat Momen, Nabeel Mohammed, and Tanzilur Rahman [9]
“Automated Status Identification of microscopic images obtained From Malaria Thin Blood Smears”. This research paper has been backed by the Ministry of Research and Technology. Equally important, the authors received support from many and specially would like to deliver the best gratitude to Andree Ang Surya and Teresa Vania Tjahja from Swiss German University for the supports and scholarly inputs.[10]
Malaria Parasite detection using different machine learning classifier watershed segmentation technique to acquire plasmodium infected and non-infected erythrocytes and relevant feature was extracted. In the studies, six different machine learning approaches for classification were applied.[11]
In this paper, a complete review and analysis of different ways to distinguish images especially in a discussion of how to stop [5], [7], [9], [16]. The goal of this study is to explore advantages and disadvantages of the thresholding of several images’ methods in the medical model [14]. Fewer locations selected Threshold methods are close to Fuzzy C-Mean Algorithm's the way, the way of the Wolf, the way of Bradley, the way of Bernsen, Triangle and Deghost's Method were tested on Malaria data set.
IV. METHODS AND MATERIALS
A. Dataset Details
Let's talk about the data we would use in our analysis. We are fortunate to have researchers at theThe National Library of Medicine's Lister Hill National Center for Biomedical Communications (LHNCBC) who carefully compiled and interpreted this database of healthy and infected blood smear images. We downloaded these databases from Kaggle.com.
To reduce the burden on microscopes in used areas and to improve diagnostic accuracy, researchers at the National Library of Medicine's Lister Hill National Center for Biomedical Communications (LHNCBC) have developed a standard operating system. Android® smartphone attached to a standard light microscope (Poostchi et al., 2018).
Giemsa-contaminated blood smear slides from 150 P. falciparum-infected and 50 healthy patients were collected and photographed at Chittagong Medical College Hospital, Bangladesh. The built-in smartphone camera has detected slide images for each small viewing area. The photos were presented in person by a slide-reading expert at the Mahidol-Oxford Tropical Medicine Research Unit in Bangkok, Thailand. Unidentified photos and annotations archived in NLM (IRB # 12972)
B. Resource Requirement (Hardware and Software)
- Programming Language: Python, HTML, CSS, JS
- Hardware: 4GB RAM (At least)
- 250 GB of Hard Disk Space (At least), 1 GB of VRAM (At least)
- Software: Windows 7 or later, Ubuntu, Mac OS X or later
- Technologies Used: Convolutional Neural Network, Tensor Flow, Google Colab, Digital Image Processing.
C. Image Processing
OpenCV is used to perform image processing. OpenCV is an editing library activity primarily intended to detect real-time computer. Digital imagery is the main data source for this research. Photographs used in this study were taken at the Lister Hill National Center for Biomedical. The communications section of the National Library for The tree. 27,203 images (both viral and non-viral) were used in model development.
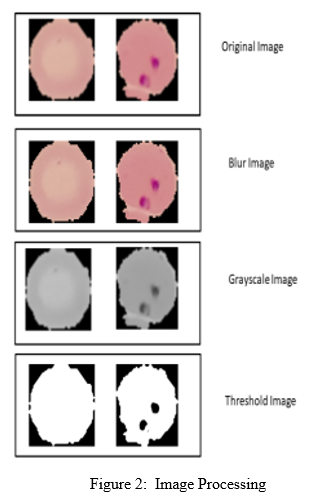
These the images are very detailed and enlarged. RBC's images taken with a small smear continued developed and analyzed in relation to the Malaria Parasite adoption. This way, instead of a box filter combining coefficients of equal filtering, Gaussian kernel is used. Gaussian filtering is very effective in removing Gaussian sound from an image. It is achieved with the assistance of internal function by calculating the breadth and length of the kernel which should be smooth and bizarre along with the general deviations in both directions.
The blurry RGB image is converted to a gray scale image. Further on the gray scale the border is used and a black and white image is obtained. By using the built-in contour function all concerts are marked and the top 5 concert venues are retrieved and saved in csv and blood cell image form used as a database in the machine learning model.
V. PURPOSE PLAN OF WORK
The eventual net performance after training will be substantially determined by the architecture of a CNN. Deep learning's primary method entails using a multi-layer network to map the input space by modifying it at hidden nodes. Through a technique known as back-propagation, the network tries to learn the best mapping of the input data through a series of transformations. The partial derivative or gradient of the input parameters is determined using the chain rule from the partial derivative of the output, given an objective function. Thus, by measuring the changes of the next layer connected to it, the changes of one layer can be computed recursively. The feed-forward and back-propagation algorithms are used to acquire a CNN model. The output of all units in each layer is computed using the feedforward propagation method, which applies an activation function that isn't linear to the weighted sum of all inputs from the previous layer for each unit. A rectified linear unit (ReLU), hyperbolic tangent (tanh), logistic function, or other activation function can be used. By adjusting the parameters/weights of each layer, backpropagation is used to train or fine-tune the deep network. When enough training data enters and spreads across the whole network, a CNN may develop a model of the incoming data using the above propagation methods. Image Classification for Malaria Using CNN Architecture, For the malaria blood smear classification job, we use a 17-layer CNN model based on the prior explanation. The network is arranged into blocks of comparable layers, with feature maps that are 1x1 in width and height and feature representations that are 256 deep at the final completely linked layer. One convolutional layer plus one ReLU layer in a sandwich form provides for improved learning.
VI. BUILDING MACHINE LEARNING MODEL
The image processing approach outlined in the previous chapter is used to produce data. The data consists of five maximum contour areas along with the status. The pandas package is used to view the data. The data is then cleaned by deleting null values and infinite values that were produced due to an error.
At random, the dataset is separated into training and testing data. The training to testing data ratio is 4:1. The model is trained using training data, whereas testing data is used to assess it. Random Forest Algorithm was utilized as the classification algorithm. Logistic Regression, Decision Tree Classifier, Random Forest Classifier, Support Vector Machine, and Bagging Classifier are the classification techniques employed.
A. Logistic Regression
When the dependent variable is binary, i.e., present or absent, it is subjected to regression analysis. It's used to explain the relationship between the dependent and independent variables and to describe data.
B. Decision Tree Classifier
This method is employed for both classification and regression purposes. A collection of training examples is split down into smaller and smaller subgroups in this technique, while an accompanying decision tree is constructed sequentially. A decision tree encompassing the training set is returned at the finish of the learning procedure.
C. Random Forest Classifier
This method employs a huge number of separate decision trees that work together to form an ensemble. Each tree in the forest produces a class prediction, and the class with the highest votes becomes the prediction of our model.
D. Support Vector Machine
The Support Vector Machine (SVM) is a supervised learning technique that plots the dataset using the independent and dependent variables. The plotted points are divided into two categories. The output of a support vector machine is a map of the sorted data with the margins between the two as wide as feasible.
E. Bagging Classifier
Bootstrap aggregating is another name for it. It's a machine learning ensemble meta-algorithm that was created to improve precision and stability of machine learning algorithms. With the aid of a bagging classifier, you may avoid the problem of overfitting and reduce variance. We discovered that when the regularization and gamma parameters in SVM were changed, the output ranged from 84 percent to 87 percent. The accuracy ranged from 10% to 87 percent when the kernel was modified.
The random forest classifier yielded the highest accuracy.
VII. PROPOSED METHODOLGY
A. Data Source
We separated the visible region of erythrocytes in actual pictures using blood smear images received from Chittagong Medical College Hospital in Bangladesh. Our data set comprises 27,578 pictures of erythrocytes, with a 1: 1 ratio of infected cells to viral cells. All photos are sorted and standardised into medium-width training images of 44* 44 pixels and three coloured channels.
B. Data Preprocessing
In MATLAB, all photos are read, scaled as appropriate, and altered before being sent into the MatConvnet toolbox. We use the eigenvalue decomposition (EVD) function in the covariance matrix to increase the area brightness and brightness and whiten the complete database before transmitting data to the CNN network for training. A common CNN's schema is shown below. The feature extractor is made up of Convolutional and max-pooling layers in the first half. The fully connected layer, which conducts non-linear transformations of the retrieved features and serves as a classifier, is the second portion.
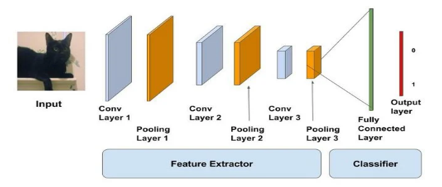
The input is fed into the network of stacked Conv, Pool, and Dense layers in the diagram above. The output can be a softmax layer that indicates whether or not there is a cat present. You may also use a sigmoid layer to determine the likelihood that the image is of a cat. Let's take a closer look at the two layers. The convolutional layer may be compared to the CNN's eyes. This layer's neurons search for certain characteristics. They create a high activation if they locate the traits they're seeking for.
In signal processing language, convolution may be conceived of as a weighted sum of two signals or functions ( in terms of mathematics ). To compute convolution at a certain position (x, y) in image processing, we extract a k x k sized chunk from the picture centred at that place (x,y). We next use the convolution filter (also sized k x k) to multiply the values in this chunk element by element, and then combine them altogether to get a single output. That concludes our discussion. The kernel size is denoted by the letter k.
C. CNN Model Training
We employ ten-fold verification throughout the data set to train and test our CNN model, with 90% of the pictures being used for training and 10% being used for testing. 90% of the photos in model training are isolated from the actual training set, with the remaining 10% utilised for back distribution validation. In addition to ten times the verification, performance test requirements include average accuracy, sensitivity, accuracy, accuracy, F1 score, and the Matthews correlation coefficient. To learn transfer, a pre-trained AlexNet based on the CIFAR-100 data set is employed as an outlet. Connected to a typical SVM separator so we can compare transfer readings to our CNN model.
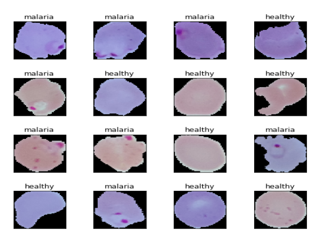
Based on the sample of the images above, we can note the subtle differences between malaria and healthy cell images. Basically, we will make our in-depth learning models try and learn these patterns during model training. We set some basic settings before we start training our models. We look forward to researching the exciting world health story of malaria detection in this article. Getting malaria on your own is not an easy task and finding the right personnel worldwide is also a major problem. We consider it easy to develop open-source strategies that use AI that can give us modern accuracy in detecting malaria and thus make AI more profitable for the community.
D. Result
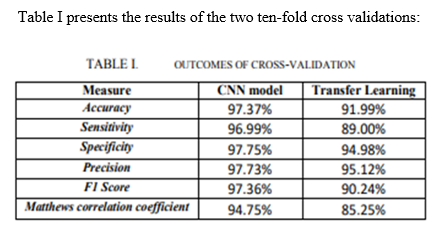
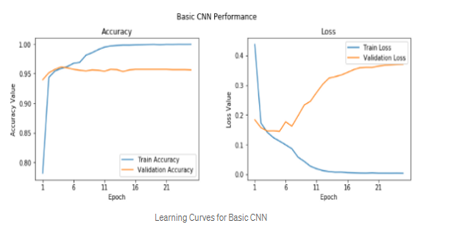
The results show that the new CNN model has much higher performance compared to the transfer learning model. CNN model class accuracy is 97.37%, and model sensitivity, specificity, and accuracy all reach 97%. The F1 school and Matthew’s coefficient (MCC) of the CNN trained model are both 7% larger than the transfer study model. This shows that the CNN trained model is a much better representation of the training images than the transfer learning model, which relies on a feature-based rendering of a pre-trained model in a completely different image set.
VIII. ACKNOWLEDGMENT
This project was completed under Dr. Vasudha Vashisht who is our main guide for this project and supported by Department of Computer Science and Engineering, Amity University Uttar Pradesh. Thanks for all the support to our Guide Dr Vasudha Vashisht and Amity University.
Conclusion
We conclude at the end of this research that our newly constructed convolutional neural network model is an excellent option for separating blood smears. After training with almost 27,000 pictures, the CNN model shows great stage performance when compared to study transfers and other similar studies [7, 8, 23]. The content and volume of the training data have an impact on its performance. Following our earlier in-depth research of genomes, we anticipate this in-depth study will considerably enhance the efficiency and accuracy of malaria diagnoses and other health-related applications.
References
[1] Mahdieh Poostchia, Kamolrat Silamutb, Richard J. Maudebcd, Stefan Jaegera, George Thoma, “Image analysis and machine learning for detecting malaria”. Received 30 October 2017, Revised 7 December 2017, Accepted 19 December 2017, Available online 12 January 2018. [2] W. A. Mustafa, R. Santiagoo, I. Jamaluddin, N. S. Othman, W. Khairunizam and M. N. K. H. Rohani, \"Comparison of Detection Method on Malaria Cell Images,\" 2018 International Conference on Computational Approach in Smart Systems Design and Applications (ICASSDA), 2018, pp. 1-6, doi: 10.1109/ICASSDA.2018.8477624. [3] Y. M. Kassim et al., \"Clustering-Based Dual Deep Learning Architecture for Detecting Red Blood Cells in Malaria Diagnostic Smears,\" in IEEE Journal of Biomedical and Health Informatics, vol. 25, no. 5, pp. 1735-1746, May 2021, doi:10.1109/JBHI.2020.3034863. [4] S. S. Savkare Moze College of Engineering, University of Pune; S. P. Narote Sinhgad College of Engineering, University of Pune, Pune, India. [5] Jane Hung, Anne Carpenter; Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) Workshops, 2017, pp. 56-61. [6] THOMAS HA¨ NSCHEID, JOSE´ MELO-CRISTINO, AND BERNADINO G. PINTO Department of Clinical Pathology, Hospital Santa Maria, Lisbon, Portugal [7] Aimon Rahman, Hasib Zunair, M Sohel Rahman, Jesia Quader Yuki, Sabyasachi Biswas, Md Ashraful Alam, Nabila Binte Alam, M.R.C. Mahdy. [8] Z. Liang et al., \"CNN-based image analysis for malaria diagnosis,\" 2016 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 2016, pp. 493-496, doi: 10.1109/BIBM.2016.7822567. [9] Deep Learning Based Automatic Malaria Parasite Detection from Blood Smear and Its Smartphone Based Application by K. M. FaizullahFuhad, Jannat Ferdousey Tuba, Md. Rabiul Ali Sarker, Sifat Momen, Nabeel Mohammed, and Tanzilur Rahman [10] Dian Anggraini, Anto Satriyo Nugroho, Christian Pratama, Ismail Ekoprayitno Rozi, Aulia Arif Iskandar, Reggio Nurtanio Hartono, Swiss German University Campus EduTown, BSD City Tangerang15339 Indonesia [11] Adedeji Olugboja, Zenghui Wang “Malaria Parasite detection using different machine learning classifier”2017 international conference of machine learning Date of Conference: 9-12 July 2017 Date Added to IEEE Xplore: 16 November 2017 DOI: 10.1109/ICMLC.2017.8107772 [12] Z. Huang and K. Chau, “A New Image Thresholding Method Based on Gaussian Mixture Model,” Appl. Math. Comput., vol. 205, no. 2, pp. 899–907, 2008. [13] A. Dirami, K. Hammouche, M. Diaf, and P. Siarry, “Fast multilevel thresholding for image segmentation through a multiphase level set method,” Signal Processing, vol. 93, no. 1, pp. 139–153, 2013. [14] H. Ahmady Phoulady, D. B. Goldgof, L. O. Hall, and P. R. Mouton, “Nucleus segmentation in histology images with hierarchical multilevel thresholding,” in Proc. SPIE 9791, Medical Imaging 2016, 2016, vol. 9791, p. 979111. [15] S. Athinarayanan, M. V Srinath, and R. Kavitha, “Computer Aided Diagnosis for Detection and Stage Identification of Cervical Cancer by Using Pap Smear Screening Test Images,” ICTACT J. Image Video Process., vol. 6, no. 4, pp. 1244–1251, 2016. [16] W. A. Mustafa, H. Yazid, S. Yaacob, and S. Basah, “Blood vessel extraction using morphological operation for diabetic retinopathy,” IEEE Reg. 10 Symp., no. 3, pp. 208–212, Apr. 2014
Copyright
Copyright © 2022 Mayank Singh, Rishabh Khurana, Parichay Jain, Ayush Verma. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET42831
Publish Date : 2022-05-17
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

