Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- References
- Copyright
Modification of DNA by Genetic Engineering
Authors: Harpreet Singh
DOI Link: https://doi.org/10.22214/ijraset.2023.51603
Certificate: View Certificate
Abstract
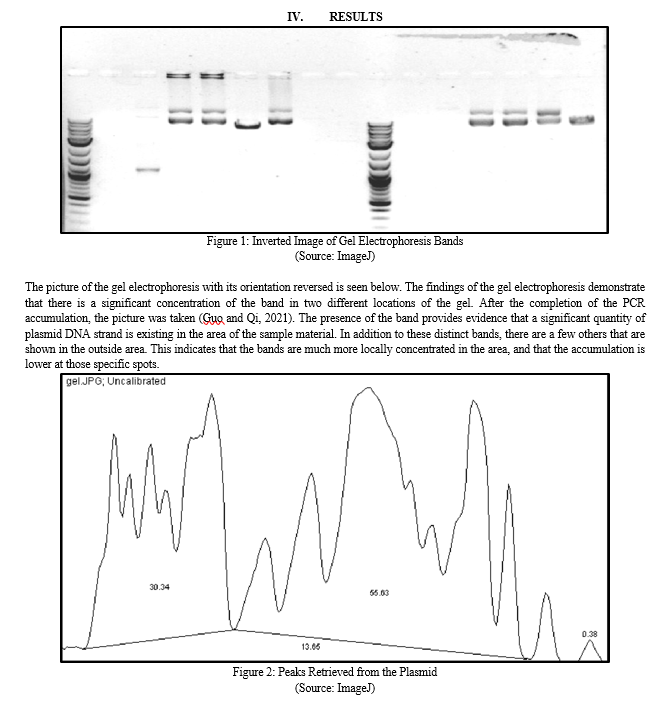
Genetic engineering modifies a living organism\'s DNA in a lab. Altering a single DNA base pair (A-T or C-G), deleting or inserting DNA, or employing all three approaches together may accomplish this purpose. Inserting a gene with a desired characteristic into a separate species\' genome is one use of genetic engineering. Genetic modification has been utilized to develop cancer drugs, brewing yeasts, GM crops and animals, and other commercial and scientific items. Probe tests on genomic DNA and RNA can identify certain sequences. Short DNA probes may be detected using radioactive or fluorescent reagents. Gel electrophoresis separates different-sized nucleic acid fragments. Blotting transfers gel particles to nylon membranes. Radioactive or fluorescent probe sequences may identify trans-membrane nucleic acid components. Southern blotting transfers DNA to a nylon membrane, whereas northern blotting transfers RNA. Northern blots identify gene expression, whereas southern blots examine nucleotide patterns. Below is a reversed gel electrophoresis image. Gel electrophoresis shows the band is concentrated in two locations. After PCR amplification, the snapshot was taken. The band suggests a substantial quantity of plasmid DNA in the sample. The outside area also has a few more distinct bands. This means the bands are localized and accumulation is lesser.
Introduction
I. INTRODUCTION
Genetic engineering or genetic manipulation refers to the process that takes place in the lab when the DNA of a living creature is altered in some way. This may be accomplished by modifying a single DNA base pair (A-T or C-G), deleting or adding DNA, or a combination of these three methods. One use of genetic modification would be to transfer a gene from one species into the genome of an organism from a different species in order to bring about the manifestation of a certain characteristic. The creation of cancer therapies, brewing yeasts, genetically modified crops and animals, and a great number of other goods with industrial and scientific uses have all been accomplished via the use of genetic modification. Artificial genetics. In the field of genetic modification, there has been a transition away from the practise of cloning organisms for the purposes of research and experimental application and toward the practise of real bioengineering in order to gain more knowledge and develop innovative biological capabilities. Students will get an understanding of the foundations of isolating and analyzing plasmid DNA using bacteria via participation in this laboratory activity. Students will learn how to utilize the Polymerase Chain Reaction (PCR) to amplify specific DNA as well as how to purify plasmids in this laboratory activity. In vitro gen Life Technologies in the United Kingdom sells the cloning host strain E. coli TOP10, which may be purchased from them (Roy et al. 2019). The plasmid DNA research and development community makes extensive use of this strain. This category is considered to have a Danger Level 1 rating. There is no information available on the insert size of the plasmid pUB1, which was produced from pET101/D-TOPO (DNA inserted at TOPO recognition sites).
In the area of recombinant DNA technology, type II restriction endo nucleases, which are also known as restriction enzymes, are a vital component. These enzymes are able to recognise and cut only certain portions of DNA, and this ability makes them an indispensable tool. Each restriction enzyme has its own unique optimum setting that, when combined with the appropriate buffering mixture and annealing temperature, yields the very best possible outcomes. In many cases, the names given to proteins are taken from the species in which they were discovered for the first time. Agarose gel electrophoresis is a common method used in the field of molecular biology that enables the viewing of DNA (Si et al. 2022). One conceivable use for this technology is the grouping of DNA fragments into categories based on size comparisons of the various pieces. A stain such as ethidium bromide is applied to the top of the gel in order to see the DNA that is contained inside the gel. When dealing with these chemicals, it is vital to utilize alternative disposal methods and wear protective clothes at all times. This is because the combination of these stains and DNA has the potential to form compounds that are potentially hazardous. In order to see the DNA complex, it is customary to make use of ethidium bromide in combination with UV light (Wu et al. 2022). This is done so that the DNA may be seen. In order to lessen the dangers that come with being exposed to ultraviolet light, it is essential to take precautions and wear protective clothing and eyewear. The principal aim of the research is to perform an in depth analysis of the plasmid DNA manipulation process by the help of electrophoresis and PCR amplification technique.
II. REVIEW OF LITERATURE
Manipulation of DNA is a rapidly evolving field that has gained immense popularity in the last few decades. The ability to manipulate DNA has opened up new avenues for research in the fields of genetics, biotechnology, medicine, and agriculture. In this literature review, we will explore the various techniques used for manipulating DNA and their applications.
One of the earliest techniques for manipulating DNA was the use of restriction enzymes. Restriction enzymes are enzymes that cut DNA at specific sites, creating fragments of DNA. This technique has been used extensively in DNA cloning and genetic engineering. For example, restriction enzymes have been used to create recombinant DNA molecules by cutting and pasting together different DNA fragments (1).
Another technique for manipulating DNA is polymerase chain reaction (PCR). PCR is a technique used to amplify DNA sequences. It involves the use of a DNA polymerase enzyme to replicate a specific DNA sequence in vitro. PCR has been used in a wide range of applications, including DNA sequencing, genetic testing, and gene expression analysis (2).
CRISPR-Cas9 is a relatively new technique for manipulating DNA that has gained a lot of attention in recent years. CRISPR-Cas9 is a gene editing tool that allows scientists to make precise changes to DNA sequences. It involves the use of a Cas9 enzyme and a guide RNA to target and cut specific DNA sequences. CRISPR-Cas9 has been used in a wide range of applications, including gene therapy, disease modeling, and agricultural biotechnology (3).
In addition to these techniques, there are several other methods for manipulating DNA, including DNA sequencing, DNA microarrays, and gene synthesis. These techniques have been used in a wide range of applications, including genetic engineering, drug discovery, and personalized medicine.
In conclusion, the ability to manipulate DNA has revolutionized the field of genetics and opened up new avenues for research in a wide range of fields. The techniques discussed in this literature review are just a few examples of the many methods available for manipulating DNA. As technology continues to advance, it is likely that new and innovative techniques for manipulating DNA will be developed, further expanding our understanding of genetics and its applications.
III. MATERIALS AND METHODS
Nucleic acids are polymers composed of nucleotides (a sugar, a phosphate, and a nitrogenous base) joined by phosphodiester bonds in order to comprehend the essential techniques used while dealing with nucleic acids. Each of these compounds has a negative phosphate group total value. The genome is the whole of the DNA molecules contained in the nucleus. The two strands of DNA are held together by hydrogen bonds between pairs of nucleotides. DNA denaturation disassembles the molecule into its component strands, which may be reassembled by cooling. DNA polymerase has the ability to replicate DNA (Keller et al.2018). RNA molecules are migratory, while DNA stays in the nucleoplasm. Since messenger RNA (mRNA) is a direct representation of the alleles that code for proteins and is thus generated in cells, it is the most often researched type of RNA.
Studying or modifying nucleic acids requires isolating DNA or RNA from cells as the primary step. There are many ways to do this. Most approaches for extracting nucleic acids include disassembling the cell and employing enzymes to destroy macromolecules that are unnecessary. To break down cells, a lysis buffer (a fluid composed mostly of surfactant) is used. The definition of "lysis" is "to divide." These enzymes are responsible for separating lipid molecules in membranes of cells and nuclei (Ma et al. 2019). To neutralize macromolecules, enzymes, such as proteases that destroy proteins and ribonucleases (RNAses) that digest RNA, are used. The DNA is then precipitated with alcohol.
To be more exact, nucleic acids exist as negatively charged ions that may be mobilized by an electric field at neutral or basic pH in water. Using this charge, gel electrophoresis is a technique for sorting molecules by size, and chromosomes may be separated either as entire units or as smaller subunits. The nucleic acids are then pulled toward the positive electrode after being loaded into a slit at the negative electrode of a porous gel matrix (Benitez et al. 2020). Smaller molecules travel through the pores of the gel more rapidly than their larger counterparts. Molecules may be measured with molecular weight reference samples to determine their size. To see nucleic acids in a gel matrix, several fluorescent or coloured dyes may be utilized. Different nucleic acid fragments appear as bands at diverse positions on the gel, at the negative electrode end, based on their sizes.
The polymerase chain reaction (PCR) is a technique for amplifying specific DNA segments. Laboratory uses of the polymerase chain reaction (PCR) include pcr amplification for sequencing, the detection of foreign DNA contamination, and the cloning of gene segments for the study of genetic diseases. Paternity testing and genetic disease detection are two more tangible applications.
Genomic DNA and RNA are two examples of nucleic acid materials that may be probe-tested for the presence of certain sequences. Probes, which are small DNA fragments, are labeled with radioactive or fluorescent dyes to aid in detection. Gel electrophoresis resolves nucleic acid fragments of differing sizes. Blotting is the transfer of particles from a gel to a nylon membrane.
Radiation or fluorescent labeling of certain probe sequences permits identification of nucleic acid components bound to the transmembrane (Berrios Adorno, 2022). In southern blotting, DNA is transferred to a nylon membrane, whereas in northern blotting, RNA is transferred to a nylon membrane. Northern blots are used to detect gene expression, whereas southern blots are used to evaluate the availability of a certain Nucleotide pattern in a chromosome.
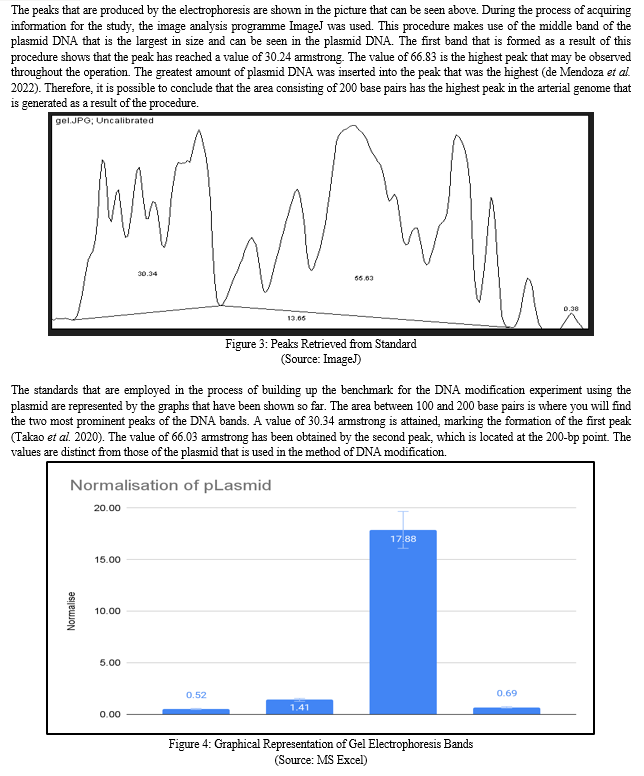
During this step of the procedure, the normalization value is determined. After dividing the region at which the peaks are obtained, we were able to arrive at the normalization value. It is quite vital to conduct an analysis of the normalized value in order to acquire information on the area of the genome in which the presence of artificial DNA fragments is more prevalent. The above graph makes it abundantly evident that the third section of the band has a rather high concentration of the synthesized DNA (Davis and Jorgensen, 2022). According to the normalized result, the amount of DNA that is present in the region is 18 times more than the amount of DNA that is present in an area that is comparable in the standard DNA sample.
V. DISCUSSION
A simple kind of genetic modification is accomplished by the use of genetic engineering to single out, duplicate, and then modify the DNA of specific cells, tissues, or species. The majority of approaches involve directly manipulating DNA in order to achieve the goal of controlling the expression of certain genes. The basic objective of genetic engineering, as is generally understood and accepted, is the direct change of DNA sequences (Cui et al. 2022). These approaches include isolating, splicing, and moving discrete pieces of DNA that correspond to particular genes in some capacity. The genetic alteration of microorganisms is made feasible by molecular genetics as well as recombinant DNA technology; nevertheless, the procedure is more difficult and costly when applied to complex organisms. Cloning and other techniques of reproductive modification, together with artificial generation processes like in vitro fertilization, embryo transfer, and artificial insemination, play an important part in the evolution of mammalian species. The current research into genetic modification of organisms is geared at a wide variety of potential applications in the fields of agriculture, pharmacology, and medicine (Cheon et al. 2019). Research on mammalian cells mostly focuses on finding ways for gene therapy, which involves swapping flawed copies of a gene for healthy copies of the gene in order to cure genetic illnesses such as cystic fibrosis. The genetically edited knockout mouse is only one example of how genetically altered animals may be used to research human genetic disorders and identify the relevance of certain sections of the genome.
Gene delivery is a technique that is used to transfer foreign DNA into host cells in order to carry out the process of genetic engineering of plants. Researchers are now using a diverse set of gene delivery methods for a variety of applications, ranging from bacterial cells to human organs. Methods are often classified into one of two categories: those that do not make use of viruses, and those that do. Because of the alterations they undergo, the viruses that are utilized in down therapeutic gene delivery techniques are unable to multiply, but they are still capable of transporting DNA for the synthesis of proteins. In order to move genetic material from one location to another, vectors such as different viruses, such as adenoviruses, retroviruses, and lentiviruses, are used (Garoli et al. 2019). Examples of non-viral procedures include electroporation, microinjection and gene gun, continuous infusion, sonication, and lipofection. Physical techniques such as "electroporation, microinjection and gene gun, continuous infusion" are also included. Gene delivery systems use a variety of tactics in order to assist the targeted cell's receipt of the genetic material that is being supplied. Understanding cytoplasmic traffic and the process of gene targeting is necessary in order to design a technique of gene delivery that is more efficient. When a pharmaceutical medicine is said to be "cell-targeted," this indicates that it is delivered specifically to a certain region or organelle inside a cell. Endocytosis is the technique of choice in the area of gene therapy for the cellular absorption of non-viral gene delivery technologies. Endocytosis is the mechanism that brings the delivery system into the cell, and subsequent cellular release is what puts in action the processes of DNA transcription and translation, which ultimately results in the production of the matching protein (Ryssy et al. 2021). A genetic delivery strategy has to eliminate any potential of an obstructing inflammatory process and overcome difficulties at each step of the process in order to maximize the amount of gene activity that can be extracted from a given organism.
References
[1] Benitez, E.K., Lomova Kaufman, A., Cervantes, L., Clark, D.N., Ayoub, P.G., Senadheera, S., Osborne, K., Sanchez, J.M., Crisostomo, R.V., Wang, X. and Reuven, N., 2020. Global and local manipulation of DNA repair mechanisms to alter site-specific gene editing outcomes in hematopoietic stem cells. Frontiers in genome editing, 2, p.601541. [2] Berrios Adorno, K.N., 2022. Engineering Controllable And Efficient Base Editors By Targeted Manipulation Of Dna Deaminases. [3] Cheon, H., Paik, J.H., Choi, M., Yang, H.J. and Son, J.H., 2019. Detection and manipulation of methylation in blood cancer DNA using terahertz radiation. Scientific reports, 9(1), pp.1-10. [4] Cui, M., Zhao, X., Reddavide, F.V., Gaillez, M.P., Heiden, S., Mannocci, L., Thompson, M. and Zhang, Y., 2022. Nonlinear manipulation and analysis of large DNA datasets. Nucleic acids research, 50(15), pp.8974-8985. [5] Roberts RJ. Restriction enzymes and their uses. Nucleic Acids Res. 1979;7(2):197-208. doi:10.1093/nar/7.2.197 [6] Mullis KB, Faloona FA. Specific synthesis of DNA in vitro via a polymerase-catalyzed chain reaction. Methods Enzymol. 1987;155:335-350. doi:10.1016/0076-6879(87)55023-6 [7] Doudna JA, Charpentier E. The new frontier of genome engineering with CRISPR-Cas9. Science. 2014;346(6213):1258096. doi:10.1126/science.1258096 [8] Davis, M.W. and Jorgensen, E.M., 2022. ApE, a plasmid editor: a freely available DNA manipulation and visualization program. Frontiers in Bioinformatics, 2. [9] de Mendoza, A., Nguyen, T.V., Ford, E., Poppe, D., Buckberry, S., Pflueger, J., Grimmer, M.R., Stolzenburg, S., Bogdanovic, O., Oshlack, A. and Farnham, P.J., 2022. Large-scale manipulation of promoter DNA methylation reveals context-specific transcriptional responses and stability. Genome biology, 23(1), pp.1-31. [10] Garoli, D., Yamazaki, H., Maccaferri, N. and Wanunu, M., 2019. Plasmonic nanopores for single-molecule detection and manipulation: toward sequencing applications. Nano letters, 19(11), pp.7553-7562. [11] Guo, A.J. and Qi, H., 2021. Using Artificial Neural Networks to Model Errors in Biochemical Manipulation of DNA Molecules. IEEE/ACM Transactions on Computational Biology and Bioinformatics, 19(5), pp.3060-3067. [12] Keller, S.M., Doherty, T.S. and Roth, T.L., 2018. Pharmacological manipulation of DNA methylation in adult female rats normalizes behavioral consequences of early-life maltreatment. Frontiers in Behavioral Neuroscience, 12, p.126. [13] Ma, F., Chen, Y., Zhu, Y. and Liu, J., 2019. Electrogenerated chemiluminescence biosensor for detection of mercury (II) ion via target-triggered manipulation of DNA three-way junctions. Talanta, 194, pp.114-118. [14] Roy, M., Chakraborty, S., Mali, K., Banerjee, A., Ghosh, K. and Chatterjee, S., 2019, August. Biomedical image security using matrix manipulation and DNA encryption. In International Ethical Hacking Conference (pp. 49-60). Springer, Singapore. [15] Ryssy, J., Natarajan, A.K., Wang, J., Lehtonen, A.J., Nguyen, M.K., Klajn, R. and Kuzyk, A., 2021. Light?Responsive Dynamic DNA?Origami?Based Plasmonic Assemblies. Angewandte Chemie, 133(11), pp.5923-5927. [16] Si, W., Zhu, Z., Wu, G., Zhang, Y., Chen, Y. and Sha, J., 2022. Encoding Manipulation of DNA?Nanoparticle Assembled Nanorobot Using Independently Charged Array Nanopores. Small Methods, p.2200318. [17] Takao, R., Shoji, T., Linklater, D., Juodkazis, S. and Tsuboi, Y., 2020 Manipulation of DNA using Nano-structured Semiconductor-assisted (NASSCA) Optical Tweezers. In Optical Manipulation and Structured Materials Conference (Vol. 11141, pp. 1114101-149). [18] Wu, S., Xu, R., Su, M., Gao, C., Liu, Y., Chen, Y., Luan, G., Jia, X. and Wang, R., 2022. A pyrF-Based Efficient Genetic Manipulation Platform in Acinetobacter baumannii To Explore the Vital DNA Components of Adaptive Immunity for IF CRISPR-Cas. Microbiology spectrum, pp.e01957-22.
Copyright
Copyright © 2023 Harpreet Singh. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET51603
Publish Date : 2023-05-05
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

