Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- Conclusion
- References
- Copyright
Molecular Profiling and Data Analysis of Lung Adenocarcinoma
Authors: Madhura Das, Uma Kumari
DOI Link: https://doi.org/10.22214/ijraset.2023.55657
Certificate: View Certificate
Abstract
Lung adenocarcinoma, a major cause of cancer-related deaths globally, was examined through a comprehensive analysis of 230 resected tumors. This study integrated various sequencing techniques to profile messenger RNA, microRNA, and DNA, along with evaluating copy number variations, methylation patterns, and proteomic data. The findings revealed a notable rate of somatic mutations (average of 8.9 mutations per megabase). Among the 18 significantly mutated genes were RIT1 activating mutations and newly identified MGA loss-of-function mutations, which were mutually exclusive with MYC amplification. Gender-based differences were observed, with EGFR mutations being more common in females and RBM10 mutations in males. Genetic abnormalities in NF1, MET, ERBB2, and RIT1 were detected in 13% of cases, often in tumors lacking other activated oncogenes, suggesting their potential driver roles. Splicing alterations were driven by somatic genomic changes, as evidenced by DNA and mRNA sequencing from the same tumor, including MET mRNA exon 14 skipping in 4% of cases. While MAPK and PI(3)K pathway activity was explained by known mutations in some cases, unexplained mechanisms appeared to activate the pathways in others. These findings lay the groundwork for understanding lung adenocarcinoma’s molecular basis and offer insights for classification and further investigations. Molecular profiling is crucial for identifying actionable mutations, guiding treatment choices in advanced adenocarcinoma and acquired resistance to tyrosine kinase inhibitors. Circulating tumor DNA sequencing gains importance in NSCLC diagnostics due to its accessibility, allowing longitudinal disease monitoring. Notably, the overall cancer death rate dropped by 27% between 1991 and 2016, resulting in around 2,629,200 fewer expected cancer deaths compared to peak death rates.
Introduction
I. INTRODUCTION
Non-small cell lung cancer (NSCLC) report as a sheet after analysis observe that for more than 80% of lung cancer cases and is the second most common cancer worldwide, with a 5-year exixtence rate of ~16%. Lung adenocarcinoma, a subtype of non-small cell lung cancer (NSCLC), has become a major global health concern, accounting for a significant portion of cancer-related deaths worldwide. Over the years, substantial progress has been made in understanding the molecular intricacies of this aggressive malignancy. This review article delves into the fascinating realm of molecular profiling and data analysis in lung adenocarcinoma, shedding light on the latest advancements that have the potential to revolutionize diagnostics, treatment strategies, and patient outcomes [1]. Lung cancer is the prime cause of cancer-related mortality and is highly resistant to conventional radiotherapy and chemotherapy are applied [2]. Using radiotherapy, there are two main types of genetic-related therapeutic strategies are targeted therapy and immunotherapy using to treat the cancer people. In Genetic mutation Targeted therapies require specific diagnosis for the patients with suffering from lung cancer. These mutations in receptors or protein kinases can affect related signaling pathways such as the RAS-RAF-MEK-ERK, PI3K-AKT-mTOR or JAK-STAT pathways, and corresponding drugs using in the targeted therapy have been developed for these targets [3]. Non-small cell lung cancer (NSCLC) is a prevalent and lethal cancer with two primary subtypes: lung squamous cell carcinoma (LUSC) and lung adenocarcinoma (LUAD) [4]. While environmental factors and genetic alterations contribute to the development and progression of LUAD, the pathogenesis of lung cancer is a multifaceted process involving complex interactions among genes, mutations, and the tumor microenvironment [5].
Current multi-omics studies revealed significant differences in the copy number variation (CNV) and methylation of LUAD subtypes with different prognoses [6], and the low expression of CNTN4 and RFTN1 estimated less successful clinical outcomes in LUAD patients [7]. Mutations in the genes EGFR, KRAS, and TP53 has a significant role in lung tumorigenesis [8]; but not all tumors develop only due to activation of these mutations and can be eliminated by suppression of these genes.Multi-omics analysis can uncover these interactions and aid in identifying molecular subtypes of LUAD, which can enhance patient prognosis through precision medicine. [9].
Lung cancer is a major cause of cancer-related deaths and is known to be resistant to traditional radiotherapy and chemotherapy [10]. To combat this, two main types of genetic-related therapeutic strategies have been developed, namely targeted therapy and immunotherapy. Targeted therapies require specific genetic mutations that are present in patients with lung cancer.
These mutations can affect related signaling pathways such as the RAS-RAF-MEK-ERK, PI3K-AKT-mTOR or JAK-STAT pathways, and drugs have been developed to target these pathways [11]. Immunotherapy involves the use of immune checkpoint blockers (ICBs) such as antibodies targeting programmed death 1 (PD-1) and cytotoxic T-lymphocyte antigen-4 (CTLA-4) but the success of this approach is heavily influenced by the tumor microenvironment (TME), including T cell abundance and tumor mutational load [12].
II. LITERATURE REVIEW
A. The Molecular Landscape of Lung Adenocarcinoma
- Genomic Alterations
The advent of next-generation sequencing (NGS) technologies has paved the way for comprehensive genomic profiling of lung adenocarcinoma. This section explores the diverse spectrum of genomic alterations that underlie the initiation and progression of this cancer. From driver mutations in genes like EGFR, KRAS, and ALK to the intricacies of tumor suppressor genes like TP53, we uncover the genetic underpinnings that shape the disease [13].
Lung adenocarcinoma is a complex and aggressive form of cancer with a high mortality rate. The emergence of next-generation sequencing (NGS) technologies has revolutionized our understanding of the genetic alterations driving this disease. Lung adenocarcinoma accounts for a significant portion of lung cancer cases and remains a major global health concern. Over the years, extensive research efforts have aimed to unravel the genetic complexities underlying this disease. The advent of NGS technologies has been a transformative milestone, enabling comprehensive genomic profiling at an unprecedented scale. One of the hallmarks of lung adenocarcinoma is the presence of driver mutations that confer a selective growth advantage to cancer cells. NGS has enabled the identification of several key driver mutations, revolutionizing our approach to targeted therapy [14]. Notably, mutations in genes such as Epidermal Growth Factor Receptor (EGFR), Kirsten Rat Sarcoma Virus (KRAS), and Anaplastic Lymphoma Kinase (ALK) have been extensively studied:
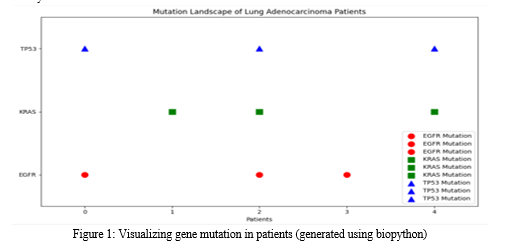
a. EGFR Mutations
EGFR mutations are among the most common driver mutations in lung adenocarcinoma. They lead to constitutive activation of the EGFR signaling pathway, promoting cell proliferation and survival. NGS has unveiled the diverse spectrum of EGFR mutations, including exon 19 deletions and the L858R substitution, guiding the development of EGFR-targeted therapies. The EGFR gene is located on the short arm of chromosome 7 (7p11.2) and encodes a 170-kDa type I transmembrane growth factor receptor with tyrosine kinase (TK) activity. It belongs to the HER/erbB family of receptor tyrosine kinases, which includes HER1 (EGFR/erbB1), HER2 (neu, erbB2), HER3 (erbB3), and HER4 (erbB4). These receptors share similar molecular structures [15].
EGFR's tyrosine kinase activity can be dysregulated by various oncogenic mechanisms, including EGFR gene mutations, increased gene copy number, and EGFR protein overexpression. Lung cancer, particularly NSCLC, is a leading cause of cancer-related mortality worldwide. Surgery is the most effective treatment for localized lung cancer, but the majority of patients present with advanced or metastatic disease, for which platinum-based chemotherapy is the standard treatment.
However, resistance to chemotherapy is common. Mutations in the TK domain of the EGFR gene were discovered in NSCLC patients, and these mutations were found to correlate with sensitivity to EGFR inhibitors like gefitinib and erlotinib. EGFR mutation status has become an important molecular predictive marker for clinical benefit from EGFR tyrosine kinase inhibitors (TKIs) [16].
b. KRAS Mutations
KRAS mutations are prevalent in lung adenocarcinoma and are associated with a more aggressive phenotype. NGS has provided insights into the heterogeneity of KRAS mutations and their potential implications for therapeutic resistance. KRAS is a downstream GTPase that can be mutated in non-small-cell lung cancer (NSCLC). Importantly, EGFR and KRAS mutations in non-small-cell lung cancer (NSCLC) appear to be mutually exclusive, meaning that a given tumor is more likely to have one or the other mutation, but not both. Despite the identification of KRAS mutations in NSCLC tumors more than two decades ago, the clinical significance of determining KRAS tumor status has only recently been appreciated. KRAS mutation status has implications for treatment decisions [17,18].
c. ALK Rearrangements
NGS has played a pivotal role in identifying ALK gene rearrangements as a driver mutation in lung adenocarcinoma. This discovery has led to the development of ALK inhibitors, such as crizotinib, as targeted therapies for a subset of patients.
While driver mutations promote cancer development, loss-of-function mutations in tumor suppressor genes contribute to unchecked tumor growth. The TP53 gene, which encodes the p53 protein, is a prominent tumor suppressor in lung adenocarcinoma. NGS has revealed the diverse landscape of TP53 mutations, their association with disease progression, and potential implications for treatment resistance. Understanding the molecular pathways that are dysregulated in lung adenocarcinoma is critical for targeted therapy development. We discuss the key signaling pathways, such as the MAPK/ERK and PI3K/AKT pathways, which play pivotal roles in tumorigenesis. Additionally, we highlight the implications of aberrant angiogenesis and immune evasion mechanisms in shaping the tumor microenvironment [19, 20].
B. Molecular Profiling Techniques
- Next-Generation Sequencing (NGS)
NGS has revolutionized cancer research by enabling the rapid and cost-effective sequencing of the entire genome, exome, or targeted gene panels. In this section, we delve into the technical aspects of NGS and its applications in lung adenocarcinoma research. From whole-genome sequencing to RNA-Seq and single-cell sequencing, we explore how these techniques provide invaluable insights into tumor heterogeneity, clonal evolution, and potential therapeutic targets. Next-generation sequencing (NGS) has catalyzed a transformative shift in cancer research, offering rapid and cost-effective genome-wide analysis. Lung adenocarcinoma, a highly heterogeneous and aggressive cancer, poses significant challenges for diagnosis and treatment. The emergence of NGS technologies has offered a promising avenue for a deeper understanding of this malignancy. The NGS process involves the creation of DNA fragment libraries followed by high-throughput sequencing. There are several NGS platforms, including Illumina HiSeq, Roche/454 Pyrosequencer, SOLiD, and Ion Torrent, each with its unique advantages and challenges. Traditional DNA sequencing methods, like Sanger sequencing, were limited in their ability to detect complex genetic alterations and were not suitable for high-throughput analysis. NGS overcomes these limitations, making it ideal for exploring the intricate genetic landscape of adenocarcinoma. NGS allows for comprehensive examination of the entire genome of adenocarcinoma cells. Researchers have identified thousands of somatic mutations, structural changes, and fusion genes within these genomes. In lung adenocarcinoma, NGS has uncovered mutation signatures associated with tobacco exposure, shedding light on the link between smoking and cancer development. NGS has revealed the presence of intratumoral heterogeneity, where different regions of the tumor have distinct genetic profiles. This insight has significant implications for treatment strategies. NGS data can be used to personalize treatment plans for adenocarcinoma patients. By identifying specific genetic alterations, clinicians can select targeted therapies that are more likely to be effective. The Cancer Genome Atlas (TCGA), an international initiative, plays a pivotal role in advancing adenocarcinoma research. By applying NGS technology to study multiple cancer types, including lung adenocarcinoma, TCGA aims to catalog genomic changes and improve cancer care[21, 22].
NGS allows for the targeted sequencing of specific genomic regions, enabling the identification of known driver mutations and other relevant alterations. Targeted sequencing panels have become invaluable tools in clinical practice for diagnosing and guiding treatment decisions in lung adenocarcinoma patients [23].
WES (whole exome sequencing) provides a comprehensive view of the exonic regions of the genome, allowing for the identification of both known and novel mutations. This approach has uncovered rare driver mutations and provided insights into the genetic heterogeneity of lung adenocarcinoma [24].
WGS (Whole Genome Sequencing) offers a complete view of the entire genome, including non-coding regions. While more resource-intensive, WGS has the potential to identify novel regulatory elements and non-coding mutations that may influence disease progression [25].
The wealth of genomic data generated by NGS has paved the way for personalized medicine in lung adenocarcinoma. By identifying specific driver mutations and their associated pathways, clinicians can tailor treatment strategies to individual patients. NGS-guided therapies, such as EGFR inhibitors and ALK inhibitors, have demonstrated improved response rates and outcomes in selected patient populations.
NGS technologies have revolutionized our understanding of the genomic landscape of lung adenocarcinoma. The identification of driver mutations, such as those in EGFR, KRAS, and ALK, along with the intricate mutational patterns of tumor suppressor genes like TP53, has deepened our knowledge of this complex disease. NGS has not only facilitated diagnosis and prognosis but has also paved the way for personalized therapeutic interventions, bringing us closer to more effective treatments and improved outcomes for lung adenocarcinoma patients. As technology continues to advance, we anticipate further discoveries that will enhance our ability to combat this challenging malignancy [22,23].
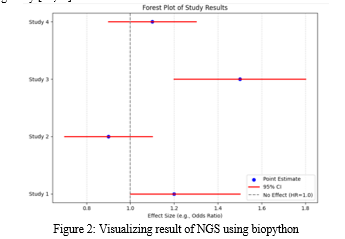
2. Liquid Biopsies
Liquid biopsies have emerged as a minimally invasive alternative to traditional tissue biopsies. We discuss the utility of circulating tumor DNA (ctDNA), circulating tumor cells (CTCs), and exosomes in monitoring disease progression, detecting minimal residual disease, and identifying actionable mutations. The promise of liquid biopsies in guiding treatment decisions is explored in depth. Adenocarcinoma is a challenging disease characterized by its diverse genetic alterations, high metastatic potential, and often late-stage diagnosis. Early detection is key to improving survival rates, and the search for less invasive and more accurate diagnostic methods has led to the development of liquid biopsy. Liquid biopsy is a non-invasive diagnostic method that involves the analysis of biomarkers present in bodily fluids such as blood, urine, or cerebrospinal fluid. These biomarkers can include circulating tumor DNA (ctDNA), circulating tumor cells (CTCs), and various proteins and nucleic acids shed by cancer cells [26,27]. Liquid biopsy offers several advantages over traditional tissue biopsies:
a. Minimally Invasive: Liquid biopsy does not require invasive procedures like surgery to obtain tissue samples. A simple blood draw or urine sample is often sufficient [28].
b. Real-Time Monitoring: Liquid biopsy allows for real-time monitoring of cancer progression and treatment response, providing valuable information for adjusting therapy [28].
c. Reduced Risk: Patients undergoing liquid biopsy experience fewer complications and risks compared to surgical biopsies, making it a safer option, especially for those with advanced-stage cancer.
d. Liquid biopsy has shown remarkable potential in the context of adenocarcinoma. Here are key ways in which it is transforming the management of this cancer:
e. Early Detection: Liquid biopsy can detect ctDNA and CTCs released by adenocarcinoma cells even in the early stages of the disease when traditional imaging may not detect tumors. This early detection can lead to more effective treatment interventions [29].
f. Treatment Selection: By analyzing the genetic mutations and alterations found in ctDNA, clinicians can tailor treatment plans to target specific pathways and mutations driving the cancer. This personalized approach enhances treatment effectiveness [29].
g. Monitoring for Recurrence: Liquid biopsy can detect minimal residual disease and monitor for cancer recurrence after surgery or treatment. Early identification of recurrence allows for timely intervention [29].
h. Assessing Treatment Response: Liquid biopsy can provide insights into how well a patient is responding to treatment. If ctDNA levels decrease, it indicates a positive response, whereas rising levels may indicate resistance or disease progression [29].
i. Identifying Resistance Mutations: Adenocarcinoma often develops resistance to targeted therapies. Liquid biopsy can identify emerging mutations that confer resistance, allowing clinicians to adjust treatment strategies promptly [29].
While liquid biopsy offers significant advantages, it is not without challenges and considerations. The sensitivity of liquid biopsy can vary, and not all mutations or biomarkers may be detectable in every patient. This requires ongoing refinement and validation of liquid biopsy techniques. Standardizing liquid biopsy protocols and analytical methods is essential to ensure consistency and reliability across different laboratories and institutions. Liquid biopsy can be costly, and its accessibility may vary depending on geographic location and healthcare infrastructure. Proper informed consent and data privacy must be upheld when using liquid biopsy for research and clinical purposes [30].
As technology and research in liquid biopsy continue to advance, the future looks promising for adenocarcinoma management. Ongoing research aims to enhance the sensitivity of liquid biopsy, making it even more effective at detecting low levels of ctDNA and CTCs. Researchers are exploring additional biomarkers beyond ctDNA and CTCs, such as microRNAs and exosomes, to provide a more comprehensive view of cancer biology. With ongoing developments and increased awareness, liquid biopsy is likely to become a standard tool in cancer diagnosis and monitoring [31].
Liquid biopsy is transforming the landscape of adenocarcinoma diagnosis and management. Its non-invasive nature, early detection capabilities, and personalized treatment insights offer new hope for patients facing this challenging disease. While challenges and considerations remain, ongoing research and technological advancements are paving the way for a brighter future in which liquid biopsy plays a central role in improving the lives of adenocarcinoma patients.
C. Data Analysis and Interpretation
- Bioinformatics Pipelines
The sheer volume of data generated by molecular profiling techniques necessitates sophisticated bioinformatics pipelines for data processing and analysis. We provide an overview of the bioinformatics tools and platforms used in lung adenocarcinoma research, emphasizing the importance of data quality control, variant calling, and integration of multi-omics data. The genomic era has ushered in an unprecedented era of data generation, particularly in the realm of lung adenocarcinoma research. The data explosion in lung adenocarcinoma research primarily stems from NGS, RNA-Seq, and other omics technologies. These techniques yield a multitude of data types, including genomic, transcriptomic, and epigenomic data. Handling such vast datasets presents numerous challenges. Issues of data quality, noise reduction, and data integration become paramount. We delve into the specific challenges associated with lung adenocarcinoma data, such as tumor heterogeneity and the need for longitudinal data to track clonal evolution. Effective data analysis begins with data quality control and preprocessing [32,33]. We outline the various steps involved in quality control, including data cleaning, adapter trimming, and base quality filtering. Mapping sequencing reads to a reference genome or transcriptome is a fundamental step in data analysis. Identifying genetic variants, including single nucleotide variants (SNVs), insertions/deletions (indels), and structural variants, is a key focus of bioinformatics pipelines. RNA-Seq data analysis encompasses gene expression quantification, differential gene expression analysis, and splicing event detection. To gain a holistic understanding of lung adenocarcinoma, researchers often integrate data from multiple omics layers, such as genomics, transcriptomics, and epigenomics. The field of bioinformatics in lung adenocarcinoma research is rapidly evolving. We speculate on the future trends, including the integration of single-cell data, the development of cloud-based analysis platforms, and the growing importance of data privacy and ethical considerations. In the era of big data, bioinformatics pipelines are the backbone of lung adenocarcinoma research. They not only enable the effective management and interpretation of complex datasets but also hold the promise of unraveling the mysteries of this heterogeneous disease. The continuous refinement of bioinformatics tools and techniques ensures that the journey of data analysis in lung adenocarcinoma research remains both challenging and rewarding, ultimately benefiting patients through improved diagnostics and personalized treatment strategies [34,35,36].
???????2. Biomarker Discovery
Identifying reliable biomarkers is a crucial step towards personalized medicine. We explore recent advancements in biomarker discovery, including the identification of novel genetic mutations, fusion genes, and immune-related markers. Additionally, we discuss the challenges of validating and translating biomarkers from research to clinical practice. Lung cancer is a deadly disease, and finding effective ways to diagnose and predict patient outcomes is crucial. Traditional methods do not account for the differences between subtypes of lung cancer, like adenocarcinoma (ADC) and squamous cell carcinoma (SCC), which have different characteristics. Computational methods, particularly deep learning and image analysis, offer a promising approach for finding valuable biomarkers that can help predict patient survival. Deep Convolutional Neural Networks (DCNNs) are used to classify different types of cells within lung cancer tissue, such as cancer cells, stromal cells, and lymphocytes. Accurate cell classification is essential for understanding how the tumor grows and spreads. The goal here is to identify the most important imaging biomarkers associated with patient survival. Feature selection methods are used to choose the most informative features while addressing the challenges posed by censoring in survival analysis. Features that are highly correlated with patients' survival outcomes are selected. Overall, the paper suggests that as computational techniques continue to advance, this approach holds great promise for improving lung cancer care and offering hope to patients battling this devastating disease [37,38,39].
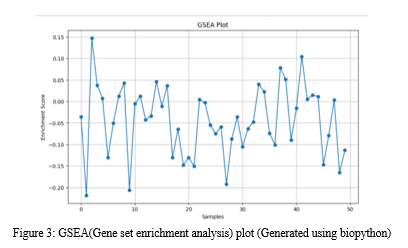
D. Clinical Implications
- Targeted Therapies
The era of precision medicine has ushered in a multitude of targeted therapies for lung adenocarcinoma. We provide an in-depth analysis of FDA-approved targeted therapies, including EGFR inhibitors, ALK inhibitors, and immune checkpoint inhibitors. Moreover, we discuss the importance of matching patients with specific genetic alterations to the most suitable treatment regimen [40].
???????2. Immunotherapy
Immunotherapy has emerged as a game-changer in the field of cancer treatment. We explore the mechanisms of immune checkpoint blockade and its role in unleashing the patient's immune system against cancer cells. The challenges and opportunities of immunotherapy in lung adenocarcinoma, including predictive biomarkers like PD-L1 expression, are thoroughly examined [41].
E. Future Directions
- Artificial Intelligence and Machine Learning
Artificial intelligence and machine learning are poised to transform the landscape of cancer research and clinical practice. We discuss how these technologies can enhance early diagnosis, predict treatment responses, and optimize treatment regimens. The potential of AI-driven drug discovery and drug repurposing is also explored.
ML models have been developed to predict the therapeutic response of immune checkpoint inhibitor (ICI) therapy for non-small cell lung cancer (NSCLC) by analyzing immune regulation-related gene profiles in patient biopsy samples. The expression of PD-L1 in tumors is influenced not only by the tumor microenvironment (TME) but also by genetic mutations, epigenetic changes (such as DNA and RNA methylation), and the presence of oncogenic and tumor suppressor signals. Accurate characterization of these factors is essential for assessing treatment efficacy. Epigenetic markers like DNA methylation (DNAm) have been used in ML algorithms to identify markers associated with cancer recurrence risk. These markers, combined with clinical data, can predict recurrence-free survival and prognosis in NSCLC patients. Acetylation, another epigenetic alteration, is being explored as a prognostic factor in early-stage lung adenocarcinoma (LUAD). Two gene signatures, RBBP7 and YEATS2, have been identified to predict recurrence-free survival [42,43,44].
Proteomics, which studies proteins and their modifications, is being used to develop predictive models for clinical outcomes and side effects of lung cancer patients. Serum proteomics has shown potential in predicting survival in lung cancer patients. Gut microbiome analysis has gained attention in recent years, and ML models have been used to predict the potential benefits of immunotherapy for NSCLC patients based on gut microbiome samples. ML algorithms have been applied to laboratory blood data, such as the neutrophil to lymphocyte ratio (NLR) and immune cell phenotypes, to predict treatment response and disease control rate in NSCLC patients receiving ICI therapy. AI is also being used to predict and understand the occurrence of immune-related adverse events (irAEs) in patients receiving immunotherapy. This includes predicting skin and cardiac irAEs, although the predictive value has been limited so far [45,46].
??????????????2. Personalized Medicine
The future of lung adenocarcinoma treatment lies in personalized medicine. We delve into the concept of patient-specific treatment strategies based on the individual's genomic profile, tumor characteristics, and immune status.
The challenges and ethical considerations associated with implementing personalized medicine on a broader scale are critically assessed [47].
Conclusion
In the realm of lung adenocarcinoma, molecular profiling and data analysis have emerged as powerful tools for unraveling the genetic complexities of the disease. This comprehensive review has journeyed through the molecular landscape, profiling techniques, data analysis methods, and clinical implications that collectively hold the promise of improving patient outcomes. As we continue to advance our understanding of this deadly malignancy, the future of lung adenocarcinoma research and treatment appears brighter than ever, offering hope to countless individuals affected by this devastating disease.
References
[1] Siegel, R. L., Miller, K. D., & Jemal, A. (2019). Cancer statistics, 2019. CA: a cancer journal for clinicians, 69(1), 7–34. https://doi.org/10.3322/caac.21551 [2] Denisenko, T. V., Budkevich, I. N., & Zhivotovsky, B. (2018). Cell death-based treatment of lung adenocarcinoma. Cell death & disease, 9(2), 117. https://doi.org/10.1038/s41419-017-0063-y [3] Heist, R. S., & Engelman, J. A. (2012). SnapShot: non-small cell lung cancer. Cancer cell, 21(3), 448.e2. https://doi.org/10.1016/j.ccr.2012.03.007 [4] Duma, N., Santana-Davila, R., & Molina, J. R. (2019). Non-Small Cell Lung Cancer: Epidemiology, Screening, Diagnosis, and Treatment. Mayo Clinic proceedings, 94(8), 1623–1640. https://doi.org/10.1016/j.mayocp.2019.01.013 [5] Vaz J, Narayan EJ, Dileep Kumar R, Thenmozhi K, Thiyagesan K, Baskaran N (2017) Prevalence and determinants of stereotypic behaviours and physiological stress among tigers and leopards in Indian zoos. PLoS ONE 12(4): e0174711. https://doi.org/10.1371/journal.pone.0174711 [6] Rong, G., Zheng, Y., Chen, Y., Zhang, Y., Zhu, P., & Sawan, M. (2023). COVID-19 Diagnostic Methods and Detection Techniques. Encyclopedia of Sensors and Biosensors, 17–32. https://doi.org/10.1016/B978-0-12-822548-6.00080-7 [7] Zhao, Y., Kuang, M., Li, J., Zhu, L., Jia, Z., Guo, X., Hu, Y., Kong, J., Yin, H., Wang, X., & You, F. (2021). SARS-CoV-2 spike protein interacts with and activates TLR41. Cell research, 31(7), 818–820. https://doi.org/10.1038/s41422-021-00495-9 [8] Kobayashi, S., Boggon, T. J., Dayaram, T., Jänne, P. A., Kocher, O., Meyerson, M., Johnson, B. E., Eck, M. J., Tenen, D. G., & Halmos, B. (2005). EGFR mutation and resistance of non-small-cell lung cancer to gefitinib. The New England journal of medicine, 352(8), 786–792. https://doi.org/10.1056/NEJMoa044238 [9] Wood, S. L., Pernemalm, M., Crosbie, P. A., & Whetton, A. D. (2014). The role of the tumor-microenvironment in lung cancer-metastasis and its relationship to potential therapeutic targets. Cancer treatment reviews, 40(4), 558–566. https://doi.org/10.1016/j.ctrv.2013.10.001 [10] Denisenko, T.V., Budkevich, I.N. & Zhivotovsky, B. Cell death-based treatment of lung adenocarcinoma. Cell Death Dis 9, 117 (2018). https://doi.org/10.1038/s41419-017-0063-y [11] Chen, J. H., Yang, J. L., Chou, C. Y., Wang, J. Y., & Hung, C. C. (2018). Indirect comparison of efficacy and safety between immune checkpoint inhibitors and antiangiogenic therapy in advanced non-small-cell lung cancer. Scientific reports, 8(1), 9686. https://doi.org/10.1038/s41598-018-27994-x [12] Binnewies, M., Roberts, E. W., Kersten, K., Chan, V., Fearon, D. F., Merad, M., Coussens, L. M., Gabrilovich, D. I., Ostrand-Rosenberg, S., Hedrick, C. C., Vonderheide, R. H., Pittet, M. J., Jain, R. K., Zou, W., Howcroft, T. K., Woodhouse, E. C., Weinberg, R. A., & Krummel, M. F. (2018). Understanding the tumor immune microenvironment (TIME) for effective therapy. Nature medicine, 24(5), 541–550. https://doi.org/10.1038/s41591-018-0014-x [13] Devarakonda, S., Morgensztern, D., & Govindan, R. (2015). Genomic alterations in lung adenocarcinoma. The lancet oncology, 16(7), e342-e351. [14] Skoulidis, F., Byers, L. A., Diao, L., Papadimitrakopoulou, V. A., Tong, P., Izzo, J., ... & Heymach, J. V. (2015). Co-occurring genomic alterations define major subsets of KRAS-mutant lung adenocarcinoma with distinct biology, immune profiles, and therapeutic vulnerabilities. Cancer discovery, 5(8), 860-877. [15] da Cunha Santos, G., Shepherd, F. A., & Tsao, M. S. (2011). EGFR mutations and lung cancer. Annual Review of Pathology: Mechanisms of Disease, 6, 49-69. [16] Niederst, M. J., Sequist, L. V., Poirier, J. T., Mermel, C. H., Lockerman, E. L., Garcia, A. R., ... & Engelman, J. A. (2015). RB loss in resistant EGFR mutant lung adenocarcinomas that transform to small-cell lung cancer. Nature communications, 6(1), 6377. [17] Karachaliou, N., Mayo, C., Costa, C., Magrí, I., Gimenez-Capitan, A., Molina-Vila, M. A., & Rosell, R. (2013). KRAS mutations in lung cancer. Clinical lung cancer, 14(3), 205-214. [18] Riely, G. J., Marks, J., & Pao, W. (2009). KRAS mutations in non–small cell lung cancer. Proceedings of the American Thoracic Society, 6(2), 201-205. [19] Paik, J. H., Choi, C. M., Kim, H., Jang, S. J., Choe, G., Kim, D. K., ... & Chung, J. H. (2012). Clinicopathologic implication of ALK rearrangement in surgically resected lung cancer: a proposal of diagnostic algorithm for ALK-rearranged adenocarcinoma. Lung cancer, 76(3), 403-409. [20] Okayama, H., Kohno, T., Ishii, Y., Shimada, Y., Shiraishi, K., Iwakawa, R., ... & Yokota, J. (2012). Identification of genes upregulated in ALK-positive and EGFR/KRAS/ALK-negative lung adenocarcinomas. Cancer research, 72(1), 100-111. [21] Dall’Olio, F. G., Conci, N., Rossi, G., Fiorentino, M., De Giglio, A., Grilli, G., ... & Ardizzoni, A. (2020). Comparison of sequential testing and next generation sequencing in advanced lung adenocarcinoma patients–a single centre experience. Lung Cancer, 149, 5-9. [22] Akazawa, Y., Saito, T., Hayashi, T., Yanai, Y., Tsuyama, S., Akaike, K., ... & Yao, T. (2018). Next-generation sequencing analysis for gastric adenocarcinoma with enteroblastic differentiation: emphasis on the relationship with hepatoid adenocarcinoma. Human pathology, 78, 79-88. [23] Drilon, A., Wang, L., Arcila, M. E., Balasubramanian, S., Greenbowe, J. R., Ross, J. S., ... & Rizvi, N. A. (2015). Broad, hybrid capture–based next-generation sequencing identifies actionable genomic alterations in lung adenocarcinomas otherwise negative for such alterations by other genomic testing approaches. Clinical cancer research, 21(16), 3631-3639. [24] Bonfiglio, S., Vanni, I., Rossella, V., Truini, A., Lazarevic, D., Dal Bello, M. G., ... & Coco, S. (2016). Performance comparison of two commercial human whole-exome capture systems on formalin-fixed paraffin-embedded lung adenocarcinoma samples. BMC cancer, 16(1), 1-18. [25] Reynolds, I. S., Thomas, V., O’Connell, E., Fichtner, M., McNamara, D. A., Kay, E. W., ... & Furney, S. J. (2020). Mucinous adenocarcinoma of the rectum: a whole genome sequencing study. Frontiers in Oncology, 10, 1682. [26] Kim, I. A., Hur, J. Y., Kim, H. J., Lee, S. E., Kim, W. S., & Lee, K. Y. (2020). Liquid biopsy using extracellular vesicle-derived DNA in lung adenocarcinoma. J. Pathol. Transl. Med, 54, 453-461. [27] Lin, L. H., Allison, D. H., Feng, Y., Jour, G., Park, K., Zhou, F., ... & Cotzia, P. (2021). Comparison of solid tissue sequencing and liquid biopsy accuracy in identification of clinically relevant gene mutations and rearrangements in lung adenocarcinomas. Modern Pathology, 34(12), 2168-2174. [28] Aguado, C., Giménez-Capitán, A., Karachaliou, N., Pérez-Rosado, A., Viteri, S., Morales-Espinosa, D., & Rosell, R. (2016). Fusion gene and splice variant analyses in liquid biopsies of lung cancer patients. Translational Lung Cancer Research, 5(5), 525. [29] Song, Z., Cai, Z., Yan, J., Shao, Y. W., & Zhang, Y. (2019). Liquid biopsies using pleural effusion-derived exosomal DNA in advanced lung adenocarcinoma. Translational Lung Cancer Research, 8(4), 392. [30] Cheung, A. H. K., Chow, C., & To, K. F. (2018). Latest development of liquid biopsy. Journal of thoracic disease, 10(Suppl 14), S1645. [31] Casula, M., Pisano, M., Paliogiannis, P., Colombino, M., Sini, M. C., Zinellu, A., ... & Palmieri, G. (2023). Comparison between Three Different Techniques for the Detection of EGFR Mutations in Liquid Biopsies of Patients with Advanced Stage Lung Adenocarcinoma. International Journal of Molecular Sciences, 24(7), 6410. [32] Kim, J., & Tan, A. C. (2012). BiNGS! SL-seq: a bioinformatics pipeline for the analysis and interpretation of deep sequencing genome-wide synthetic lethal screen. Next Generation Microarray Bioinformatics: Methods and Protocols, 389-398. [33] Lim, S. B., Tan, S. J., Lim, W. T., & Lim, C. T. (2018). A merged lung cancer transcriptome dataset for clinical predictive modeling. Scientific data, 5(1), 1-8. [34] Merino, D. M., McShane, L. M., Fabrizio, D., Funari, V., Chen, S. J., White, J. R., ... & Allen, J. (2020). Establishing guidelines to harmonize tumor mutational burden (TMB): in silico assessment of variation in TMB quantification across diagnostic platforms: phase I of the Friends of Cancer Research TMB Harmonization Project. Journal for immunotherapy of cancer, 8(1). [35] Chang, H., Sasson, A., Srinivasan, S., Golhar, R., Greenawalt, D. M., Geese, W. J., ... & Szustakowski, J. (2019). Bioinformatic methods and bridging of assay results for reliable tumor mutational burden assessment in non-small-cell lung cancer. Molecular diagnosis & therapy, 23, 507-520. [36] Keller, A., Backes, C., Al-Awadhi, M., Gerasch, A., Küntzer, J., Kohlbacher, O., ... & Lenhof, H. P. (2008). GeneTrailExpress: a web-based pipeline for the statistical evaluation of microarray experiments. BMC bioinformatics, 9, 1-6. [37] Yao, J., Wang, S., Zhu, X., & Huang, J. (2016). Imaging biomarker discovery for lung cancer survival prediction. In Medical Image Computing and Computer-Assisted Intervention–MICCAI 2016: 19th International Conference, Athens, Greece, October 17-21, 2016, Proceedings, Part II 19 (pp. 649-657). Springer International Publishing. [38] Fujii, K., Nakamura, H., & Nishimura, T. (2017). Recent mass spectrometry-based proteomics for biomarker discovery in lung cancer, COPD, and asthma. Expert Review of Proteomics, 14(4), 373-386. [39] Shao, Y., Liang, B., Long, F., & Jiang, S. J. (2017). Diagnostic microRNA biomarker discovery for non-small-cell lung cancer adenocarcinoma by integrative bioinformatics analysis. BioMed research international, 2017. [40] Ladanyi, M., & Pao, W. (2008). Lung adenocarcinoma: guiding EGFR-targeted therapy and beyond. Modern pathology, 21(2), S16-S22. [41] Saito, M., Suzuki, H., Kono, K., Takenoshita, S., & Kohno, T. (2018). Treatment of lung adenocarcinoma by molecular-targeted therapy and immunotherapy. Surgery today, 48, 1-8. [42] Gao, Q., Yang, L., Lu, M., Jin, R., Ye, H., & Ma, T. (2023). The artificial intelligence and machine learning in lung cancer immunotherapy. Journal of Hematology & Oncology, 16(1), 1-18. [43] Wang, S., Yang, D. M., Rong, R., Zhan, X., Fujimoto, J., Liu, H., ... & Xiao, G. (2019). Artificial intelligence in lung cancer pathology image analysis. Cancers, 11(11), 1673. [44] Rabbani, M., Kanevsky, J., Kafi, K., Chandelier, F., & Giles, F. J. (2018). Role of artificial intelligence in the care of patients with nonsmall cell lung cancer. European journal of clinical investigation, 48(4), e12901. [45] Huang, S., Yang, J., Shen, N., Xu, Q., & Zhao, Q. (2023, January). Artificial intelligence in lung cancer diagnosis and prognosis: Current application and future perspective. In Seminars in Cancer Biology. Academic Press. [46] Zhang, K., & Chen, K. (2022). Artificial intelligence: opportunities in lung cancer. Current Opinion in Oncology, 34(1), 44-53. [47] Seguin, L., Durandy, M., & Feral, C. C. (2022). Lung adenocarcinoma tumor origin: a guide for personalized medicine. Cancers, 14(7), 1759.
Copyright
Copyright © 2023 Madhura Das, Uma Kumari. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET55657
Publish Date : 2023-09-07
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

