Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- Conclusion
- References
- Copyright
Securing Aggregate Queries for DNA Databases
Authors: Mrs. Vanithamani, J. M. Yashwanthan, K. Jai Ganesh, G. Keerthi Vasan
DOI Link: https://doi.org/10.22214/ijraset.2023.52009
Certificate: View Certificate
Abstract
This paper addresses the problem of sharing person-specific genomic sequences without violating the privacy of their data subjects to support large- scale biomedical research projects. The proposed method builds on the framework but extends the results in a number of ways. One improvement is that our scheme is deterministic, with zero probability of a wrong answer. We also provide a new operating point in the space-time tradeoff, by offering a scheme that is twice as fast as theirs but uses twice the storage space. This point is motivated by the fact that storage is cheaper than computation in current cloud computing pricing plans. Moreover, our encoding of the data makes it possible for us to handle a richer set of queries than exact matching between the query and each sequence of the database, including: (i) counting the number of matches between the query symbols and a sequence; (ii) logical OR matches where a query symbol is allowed to match a subset of the alphabet thereby making it possible to handle a \"not equal to\" requirement for a query symbol (iii) support f for the extended alphabet of nucleotide base codes that encompasses ambiguities in DNA sequences (iv) queries that specify the number of occurrences of each kind of symbol in the specified sequence positions (v) a threshold query whose answer is \'yes\' if the number of matches exceeds a query-specified threshold. (vi) For all query types we can hide the answers from the decrypting server, so that only the client learns the answer. (vii) In all cases, the client deterministically learns only the query\'s answer, except for query type (v) where we quantify the (very small) statistical leakage to the client of the actual count.
Introduction
I. INTRODUCTION
This paper watches out for the issue of sharing individual specific genomic progressions without slighting the assurance of their data subjects to help broad scale biomedical research wanders. The proposed procedure develops the framework. However, extends the results in different ways. One change is that our arrangement is deterministic, with zero probability of a wrong answer (instead of a low probability). We in like manner give another working point in the space-time tradeoff, by offering an arrangement that is twice as brisk as theirs however uses twofold the storage space. This point is impelled by how limit is more affordable than figuring in current circulated processing evaluating plans. Likewise, our encoding of the data makes it plausible for us to manage a wealthier plan of inquiries than revise organizing between the request and each gathering of the database, including: (i) counting the amount of matches between the inquiry pictures and a progression; (ii) lucid OR matches where a request picture is allowed to facilitate a subset of the letter set thusly making it possible to manage (as a remarkable case) a "not comparable to" need for a request picture (e.g., "not a G"); (iii) support for the widened letter set of nucleotide base codes that encompasses ambiguities in DNA groupings (this happens on the DNA course of action side as opposed to the request side); (iv) request that show the amount of occasions of each kind of picture in the predefined progression positions (e.g., two "An" and four "C" and one "G" and three 'T', occurring in any demand in the inquiry decided progression positions); (v) an edge request whose answer is "yes" if the amount of matches outperforms a request decided breaking point (e.g., "no less than 7 arranges out of the 15 request demonstrated positions"). (vi) For all inquiry sorts we can disguise the fitting reactions from the interpreting server, with the objective that solitary the client takes in the proper reaction. (vii) In all cases, the client deterministically adjusts only the inquiry's answer, beside request sort (v) where we measure the (little) truthful spillage to the client of the genuine count.
II. LITERATURE SURVEY
A. Scalable and Secure Sharing of Personal Health Records in Cloud Computing
Author: M. Li, S. Yu, Y. Zheng, K. Ren, and W. Lou.
Abstract: Personal health record (PHR) is an emerging patient-centric model of health information exchange, which is often outsourced to be stored at a third party, such as cloud providers. In this paper, we propose a novel patient-centric framework and a suite of mechanisms for data access control to PHRS stored in semi trusted servers. To achieve fine-grained and scalable data access control for PHRS, we leverage attribute-based encryption (ABE) techniques to encrypt each patient's PHR file. A high degree of patient privacy is guaranteed simultaneously by exploiting multiauthority ABE.
Our scheme also enables dynamic modification of access policies or file attributes, supports efficient on-demand user/attribute revocation and break-glass access under emergency scenarios. Extensive analytical and experimental results are presented which show the security, scalability, and efficiency of our proposed scheme.
B. An SMDP-Based Service Model for Inter domain Resource Allocation in Mobile Cloud Networks
Author: H. Liang, L. X. Cai, D. Huang, X. Shen and D. Peng
Abstract: Mobile cloud computing is a promising technique that shifts the data and computing service modules from individual devices to geographically distributed cloud service architecture. In this paper, we propose a service decision making system for inter domain service transfer to balance the computation loads among multiple cloud domains.
To this end, we formulate the service request decision making process as a semi-Markov decision process. The optimal service transfer decisions are obtained by jointly considering the system incomes and expenses. Extensive simulation results show that the proposed decision making system can significantly improve the system rewards and decrease service disruptions compared with the greedy approach.
III. OBJECTIVES
There is no universal method to create a protocol for secure multi-party computation and handling aggregate queries on encrypted data is not an exception. Several holomorphic systems only support a subset of mathematical operations, like addition, or exclusive From a security perspective, only the additive and the multiplicative are classified to be IND-CPA (stands for in distinguishability under chosen plaintext attack). Partially holomorphic cryptosystems are more desirable from a performance point of view than somewhat holomorphic cryptosystems, which support a limited operation depth. Fully holomorphicsystems have a huge cost and cannot be deployed in practice.
IV. METHODOLOGY
A. Problem Statement
- Proposed System
This paper provides a new method that addresses a larger set of problems and provides a faster query response time than the technique introduced. Our approach is based on the fact that, given current pricing plans at many cloud services providers, storage is cheaper than computing. Therefore, we favor storage over computing resources to optimize cost. Moreover, from a user experience point of view, response time is the most tangible indicator of performance; hence it is natural to aim at reducing it. Our method enhances the state of the art at both the conceptual level and the implementation level. Moreover, our encoding of the data makes it possible for us to handle a richer set of queries than exact matching between the query and each sequence of the database, including.
2. Advantages Of Proposed System
a. Counting the number of matches between thequery symbols and a sequence.
b. Logical OR matches where a query symbol is allowed to match a subset of the alphabet thereby making it possible to handle (as a special case) a "not equal to" requirement for a query symbol.
c. Support for the extended alphabet of nucleotide base codes that encompasses ambiguities in DNA sequences.
d. Queries that specify the number of occurrences of each kind of symbol in the specified sequence positions.
e. A threshold query whose answer is 'yes' if the number of matches exceeds a query-specified threshold.
V. MODULES
A. Privacy Preserving
Hospitals want to protect the confidentiality of the DNA sequences that they own and no external party has the right to access these DNA sequences for privacy reasons. Thus, other parties (be it the server or the clients) should only work on encrypted sequences and never have access to the DNA. In this, modules the file which is stored by the hospital will be encrypted and then stored in clouds.
B. Secure Outsourcing
The encrypted file will be outsourced to the clouds. This solution aims not only to provide confidentiality and access controllability of outsourced data with strong cryptographic guarantee, but, more importantly, to fulfill specific security requirements from different cloud services with effective systematic way.
C. Aggregate Queries
In this modules, important queries have often in the form of how many records contain a diagnosis of disease and gene variant. Secure outsourcing of the database and allowing such type of queries without requiring the server to decrypt the data. In this hospital will set the DNA by a large sequence of characters from the alphabet representing the four nucleotide types. This alphabet can be aggregate with additional characters representing augmented in the sequence.
D. Sequence Testing
In this modules, the queries on DNA need to take into account various errors such as irrelevant mutations, incomplete specifications and sequencing errors. Clients are authorized entities in which they are allowed to perform queries on the encrypted DNA sequences.
E. Set Match Query
In this modules, we will authenticate that the query which is asked by the researcher match with the query which is given by the cloud. The hospital will set the alphabetical sequence of DNA, and the same the Alphabetic sequence have to be given by the researchers.
F. Hiding From The Decrypted Server
In this modules, the hospital will store the encrypted file to the cloud. The cloud will internally make cloud as a key holder and cloud2 has a data holder. In which every time the researcher will query the file initially the cloudl will return the key and if it matches with the hospital secret key then cloud2 will return the decrypted data.
VI. BLOCK DIAGRAM
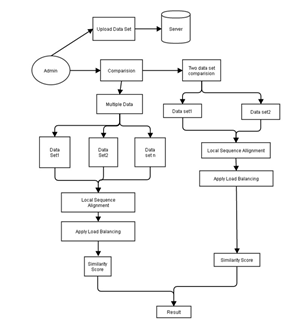
VII. ALGORITHM USED
A. AES Algorithm
AES (acronym of Advanced Encryption Standard) is a symmetric encryption algorithm. The algorithm was developed by two Belgian cryptographer Joan Daemen and Vincent Rijmen. AES was designed to be efficient in both hardware and software, and supports a block length of 128 bits and key lengths of 128, 192, and 256 bits.
B. Smith Waterman Algorithm (DNA)
The Smith-Waterman algorithm performs local sequence alignment; that is, for determining similar regions between two strings of nucleic acid sequences or protein sequences. Instead of looking at the entire sequence, the Smith-Waterman algorithm compares segments of all possible lengths and optimizes the similarity measure. In recent years, genome projects conducted on a variety of organisms generated massive amounts of sequence data for genes and proteins, which requires computational analysis. Sequence alignment shows the relations between genes or between proteins, leading to a better understanding of their homology and functionality. Sequence alignment can also reveal conserved domains and motifs.
Conclusion
We have revisited the challenge of sharing person- specific genomic sequences without violating the privacy of their data subjects in order to support large-scale biomedical research projects. We have used. the framework based on additive homomorphism encryption, and two servers: one holding the keys and one storing the encrypted records. The proposed method offers two new operating points in the space-time trade off and handles new types of queries that are not supported in earlier work. Furthermore, the method provides support for extended alphabet of nucleotides which is a practical and critical requirement for biomedical researchers. Big data analytics over genetic data is a good future work direction. There are rapid recent advancements that address performance limitations of holomorphic encryption techniques. We hope that these advancements will lead to more practical solutions in the future that can handle larger-scale genetics data. It is worth mentioning that our approach is not restricted to a fixed holomorphic encryption technique and therefore, it would be possible to use and inherit the advantages of newly developed ones.
References
[1] M. Kantarcioglu, W. Jiang, Y. Liu, and B. Malin, \"A cryptographic approach to securely share and query genomic sequences,\" Inf. Technol. Biomed. IEEE Trans., vol. 12, no. 5, pp. 606-617, 2008. [2] B. Malin and L. Sweeney, \"How (not) to protect genomic data privacy in a distributed network: using trail re-identification to evaluate and design anonymity protection systems,\" J. Biomed. Inform., vol. 37, no. 3, pp. 179-192, 2004. [3] Z. Lin, A. B. Owen, and R. B. Altman, \"Genomic research and human subject privacy,\" Science (80-. )., vol. 305, no. 5681, p. 183, 2004. [4] A. E. Nergiz, C. Clifton, and Q. M. Malluhi, \"Updating outsourced anatomized private databases,\" in Proceedings of the 16th International Conference on Extending Database Technology, 2013, pp. 179-190. [5] L. Sweeney, A. Abu, and J. Winn, \"Identifying Participants in the Personal Genome Project by Name,\" Available SSRN 2257732, 2013. [6] E. Aguiar, Y. Zhang, and M. Blanton, \"An Overview of Issues and Recent Developments in Cloud Computing and Storage Security,\" in High Performance Cloud Auditing andApplications, 2014, pp. 3-33. [7] P. Bohannon, M. Jakobsson, and S. Srikwan, \"Cryptographic Approaches to Privacy in Forensic DNA Databases,\" in Public Key Cryptography, vol. 1751, H. Imai and Y. Zheng, Eds. Springer Berlin Heidelberg, 2000, pp. 373-390. [8] F. Esponda, E. S. Ackley, P. Helman, H. Jia, and S. Forrest, \"Protecting data privacy through hard-toreverse negative databases,\" Int. J. Inf. Secur., vol. 6, no. 6, pp. 403-415, 2007. [9] F. Bruekers, S. Katzenbeisser, K. Kursawe, and P. Tuyls, \"Privacy-preserving matching of dna profiles,\" IACR Cryptol. ePrint Arch., vol. 2008, p. 203, 2008. [10] M. J. Atallah and J. Li, \"Secure outsourcing of sequence comparisons,\" Int. J. Inf. Secur., vol.4, no.4, pp. 277-287, Mar. 2005. [11] M. Blanton, M. M. J. Atallah, K. B. K.Frikken, and Q.Malluhi,\"SecureandEfficient Outsourcing ofSequence Comparisons,\" Comput. Secur. 2012, pp. 505-522,
Copyright
Copyright © 2023 Mrs. Vanithamani, J. M. Yashwanthan, K. Jai Ganesh, G. Keerthi Vasan. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET52009
Publish Date : 2023-05-11
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

