Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- Conclusion
- References
- Copyright
Similarity Coefficient Analysis using UPGMA based on Polymorphism of Isozyme and DNA Markers in Cotton
Authors: Dr. Vaibhav V. Ujjainkar
DOI Link: https://doi.org/10.22214/ijraset.2022.45362
Certificate: View Certificate
Abstract
Cotton refers to those species of genus Gossypium that bears the spinnable seed coat fibres. The proteins and enzymes are the primary product of the genes and hence are most suitable for determination of genetic purity and polymorphism. The biochemical and molecular markers may complement the efforts to reduce the burden to a greater extent, as these are estimates devoid of environmental factor which a cause of error. Especially in the selection of parents based on biochemical and molecular polymorphism, there is marked improvement in the screens for the quantitative traits and understanding their architecture. The banding pattern of biochemical markers revealed that the maximum number of bands were recorded in esterase analysis (fifteen) followed by protein analysis (twelve) whereas, only five bands each were detected in peroxidase and polyphenol oxidase analysis indicating limited polymorphism. The Relative Mobility (Rm) values were ranged from 0.083 to 0.883 (protein), 0.100 to 0.971 (esterase), 0.033 to 0.283 (peroxidase) and 0.048 to 0.206 (polyphenol oxidase).
Introduction
I. INTRODUCTION
Cotton, the white gold is one of the leading fibre crops of the country and good number of high yielding varieties and hybrids has been bred by the breeders in India. However, the requirement of better yielding lines is yet to satisfy, and therefore it is a continuous process. The conventional breeding techniques has upto last couple of decades was only a tool for improvement [14].During the recent years it has become imperative to use the modern techniques and genetic methods to fasten the process of improvement. Mahalanobis D2 Statistics is one of the powerful statistical tools for estimating the genetic distances between the germplasm [5&6]. These may be utilized for selecting parents in crossing program. Beside D2 statistics the RAPD markers can also be used to predict the genetic distance. Isozymes are form of enzymes exhibiting the same catalytic activity but differing in charge and electrophoretic mobility. The variation in banding pattern obtained between individual samples can be used to sort out genetically diverse or unrelated species which can be utilized for further improvement program.
II. MATERIAL AND METHODS
- Plant Material:The experimental material comprising of thirty-four genetically diverse and elite upland cotton lines were used for present investigation. Single seed is used for protein and DNA analysis, while the one week old seedlings are used for isozyme analysis.
- Gel Analysis:Sodium Dodecyl Sulphate Polyacrylamide Gel Electrophoresis (SDS-PAGE) method was followed for the separation of total seed proteins for the identification of cultivars [1].Alkaline Polyacrylamide Gel Electrophoresis (PAGE) followed by staining was adopted for identifying isozyme bands in different cotton cultivars [7]. Whereas, the Random Amplified Polymorphic DNA (RAPD) analysis of thirty four cotton cultivar [4].
- Data Analysis:The protein and isozyme banding pattern were analyzed by calculation of relative mobility (Rm) of band [11]

Similarity was calculated with SIMQUAL function of NTSYS that computes a variety of similarity and dissimilarity coefficients for qualitative data. The pair-wise similarity (Sij) between the accessions (i,j) were estimated using Jaccard similarity coefficients [10].
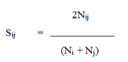
Where,
Sij = the proportion of bands common between accessions i and j
Nij =number of bands common in accession i and j
Ni = total number of bands in accession i
Nj = total number of bands in accession j
Based on genetic distance estimates, the dendrogram was constructed based on the Sij values using the clustering technique of uniweighted pair group method of arithmetic means (UPGMA) given by Nei and Li, 1979 [10].
III. RESULTS AND DISCUSSION
The proteins and enzymes are the primary products of genes and hence are most suited for genetic determination [12]. The change in coding base sequence results in corresponding replacement in amino acid and thus in primary structure of protein and enzymes. They possess ionazable groups and can therefore be made to exist in solution as electrically charged particles either as cations (+) or anions (–). The primary structure of protein and enzymes [2] in presence of a iso – electric field and while passing through a semi – porous gel medium cause dissimilar forms of protein and isoenzymes[8].
A. Banding Pattern
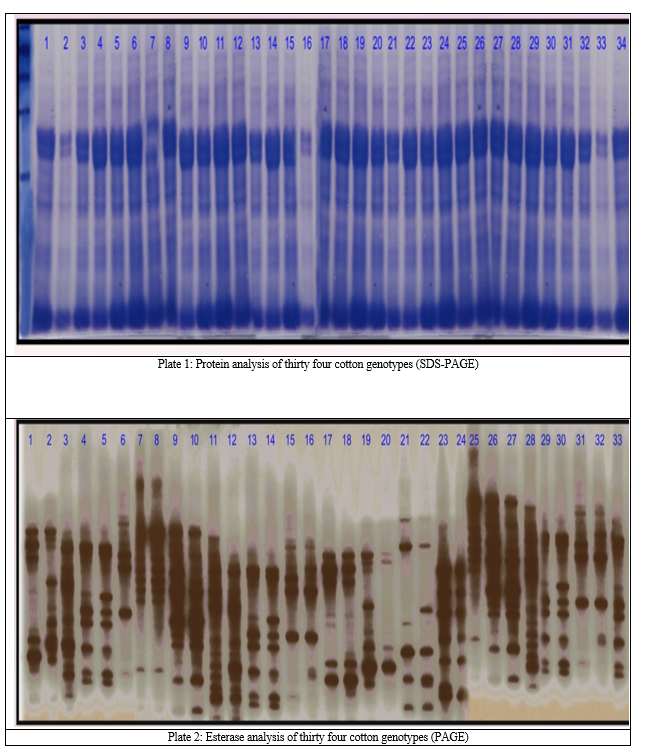
The total twelve bands were observed having relative mobility ranging from 0.083 to 0.883. The banding pattern revealed that the genotypes AKH – 62, GA – B, AKH – 023, IC – 1547 and AKH – 3436 exhibited twelve bands in the present study. Whereas, the genotypes AKH – 8835, JK – 119 and AKH – 8660 showed the least number of bands i.e. only six. The seed protein expression is usually controlled by homologous multigene families exhibiting monogenic segregation with co – dominance for molecular weight variants and presence of polypeptide bands being completely dominant over case of absence. This kind of variation, revealed that in the electrophoretic banding pattern could lead to the detection of genotype specific bands. Therefore, the polypeptides varying from their presence could be used as reliable biochemical marker for distinguishing the population. This fact confirms the congruence between the protein and genetic constitution of the individual. Beside this the number of bands can be utilized as the measure of classification but not the exact, while scoring on the basis of the presence or absence of band with specific Rm value, bears more precision than earlier one[13]. The intensity may not be the criterion for the categorization as there are many factors that may interact with the banding intensity. The migration velocity ostensibly provides the rational estimate of the degree of homology based on protein bands within and between related one amendable to statistical distance [3]. The similarity indices among the intra–cluster ranged from 0.73 to 1.00. (Table 1, Plate 1)
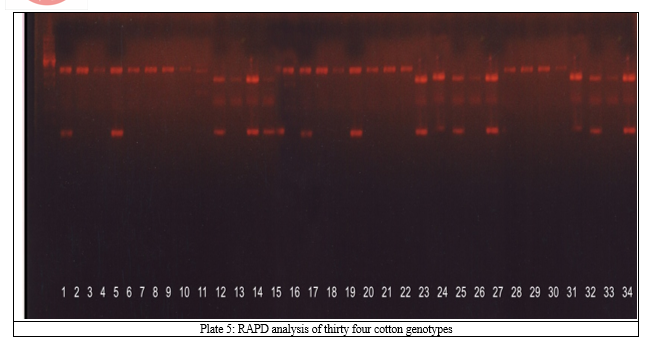
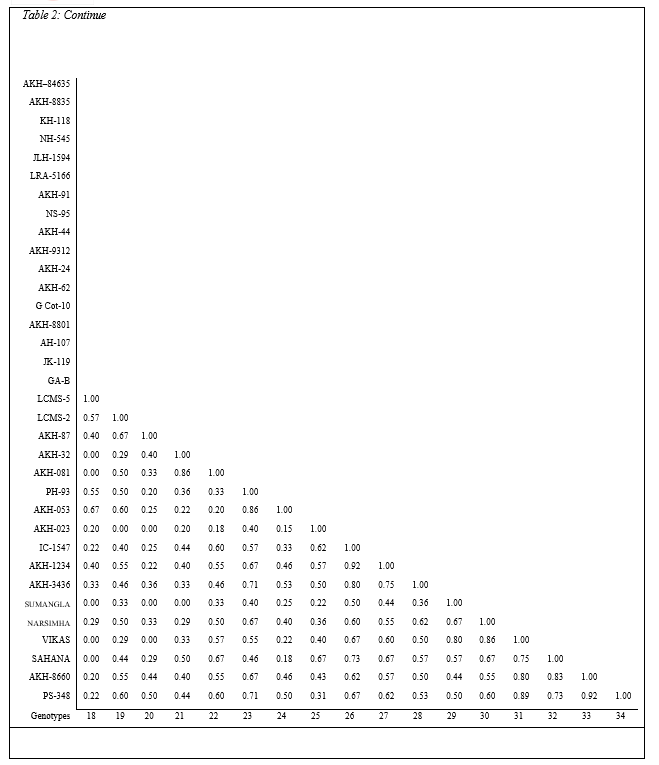
Esterases are present ubiquitously as isozymes the molecularly distinct patterns of the same enzymes. Understanding polymorphism at the enzyme level is basic to its use in population and genetic studies. The isozyme-banding pattern can be used as markers to the desirable agronomic and quality traits of different germplasm lines. Total fifteen bands were observed with range of 0.100 to 0.971 for relative mobility. The genotypes AKH – 24 and PH – 93 exhibited maximum bands (eight), followed by NH – 545, AKH – 62, AKH – 023 and AKH – 1234 of seven each. The banding pattern confirms substantial polymorphism among the genotypes tested in present conditions (Table 2, Plate 2). During the period of investigation it has been found that esterases possess very sensitive nature of enzymatic reactions, hence the results were not reproducible. Also there is limited scope for screening as they are more prone to fluctuations.
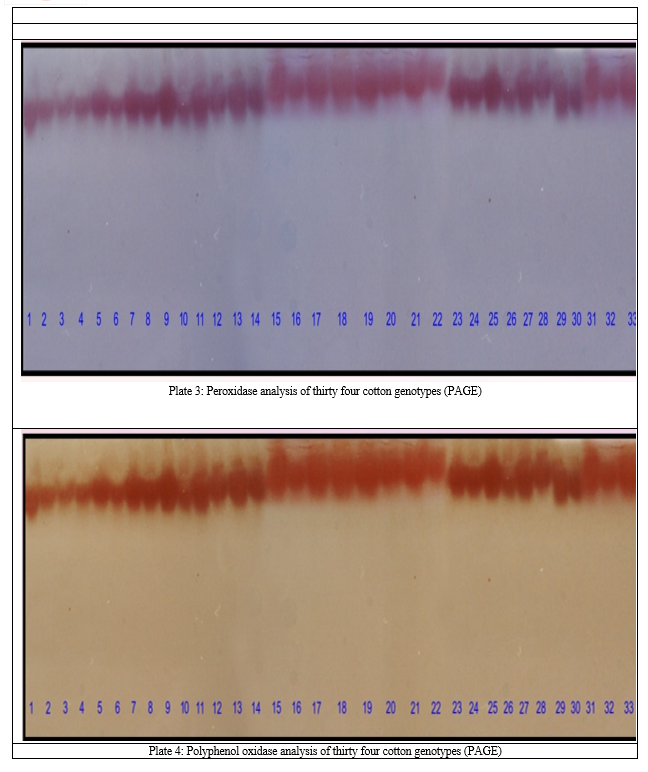
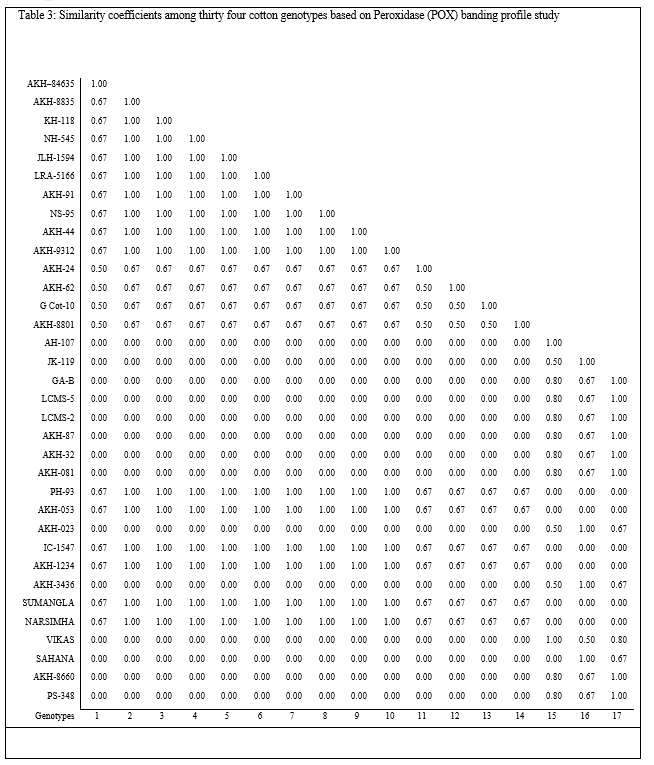
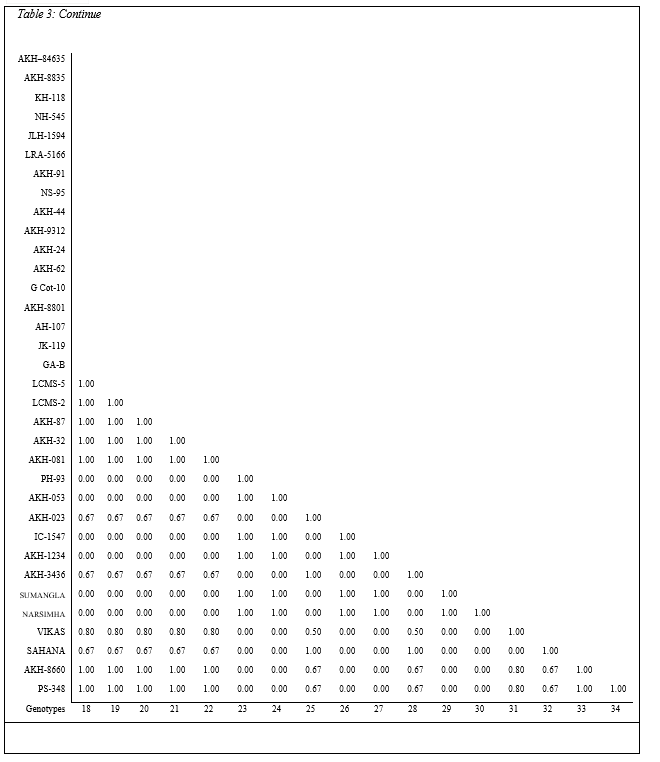
The results revealed that total five bands of peroxidase were observed in thirty four cotton lines. Very narrow range of relative mobility values, from 0.033 to 0.283 was observed, indicating that the differentiation of POX activity was not clear in cotton lines under investigation (Table 3, Plate 3).The POX possibly contributes to resistance by catalyzing oxidative polymerization of simple phenols to lignin and synthesis of antimicrobial oxidized phenols [9]. An increased peroxidases activity in various plant host parasite interactions has been correlated with the disease resistance.
Hence on the basis findings of the present study it can be revealed that the genotypes LRA – 5166, JLH – 1594, AKH – 24 and LCMS – 2 may have resistance power against pathogenic reactions on the basis of Peroxidase activity, as a factor of antibiosis kind of mechanism for plant disease resistance.
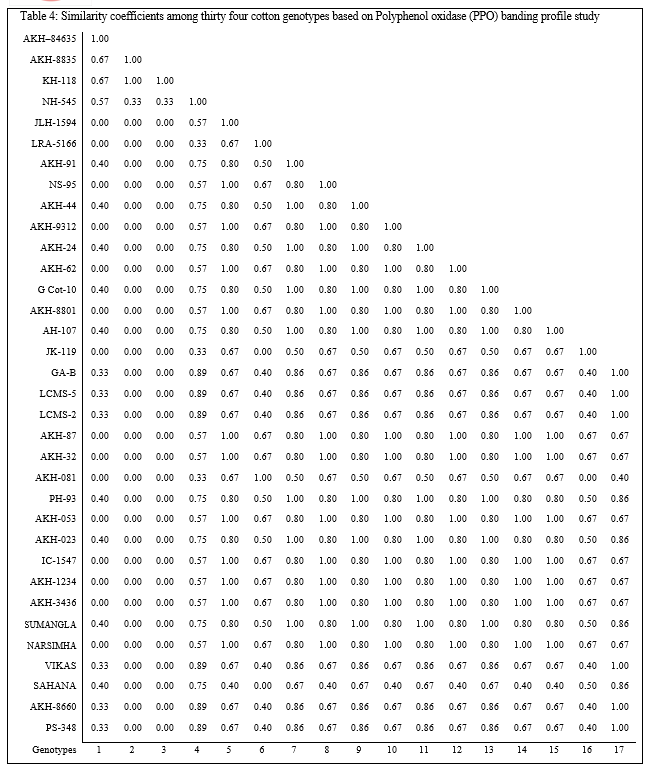
The maximum three bands were observed in AH–107 and Vikas, whereas in most of the genotypes exhibited single band of different relative mobility. Limited polymorphism was observed for banding pattern of POX for cotton lines analyzed. Total five PPO bands overlapping each other were detected having the relative mobility in the range of 0.04 to 0.21. It reveals that there is limited polymorphism within the material under study for polyphenol oxidase activity. Among the thirty four genotype, the genotype GA–B, LCMS–5, LCMS–2, Vikas, AKH–8660 and PH–348 has shown all the five bands where as the genotypes viz., AKH–8835, KH–118, NH–545, LRA–5166, JK–119 and AKH–081 exhibited single band each of varying relative mobility (Table 4, Plate 4).
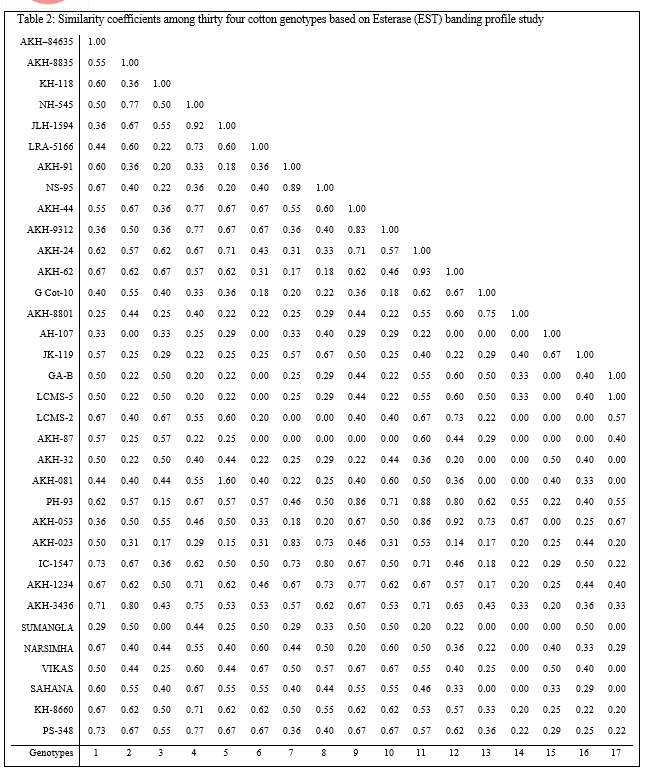
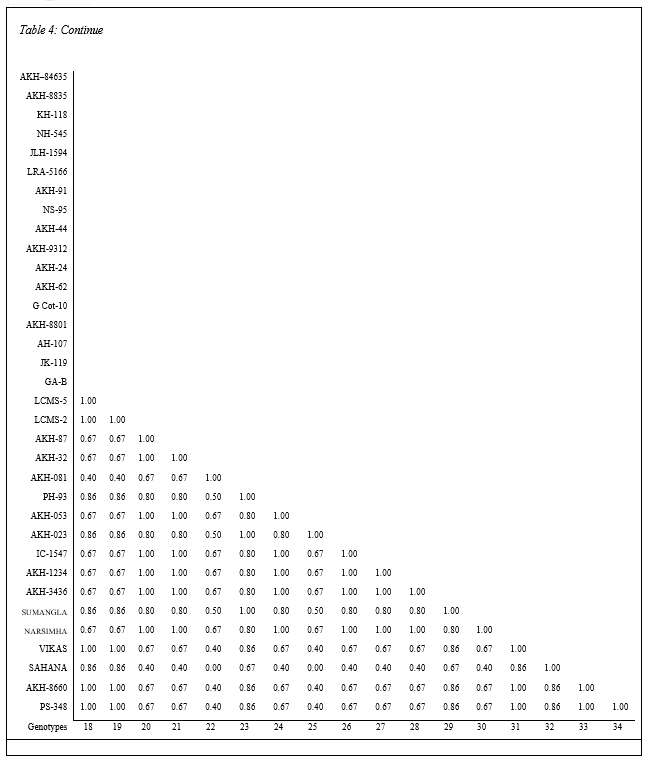
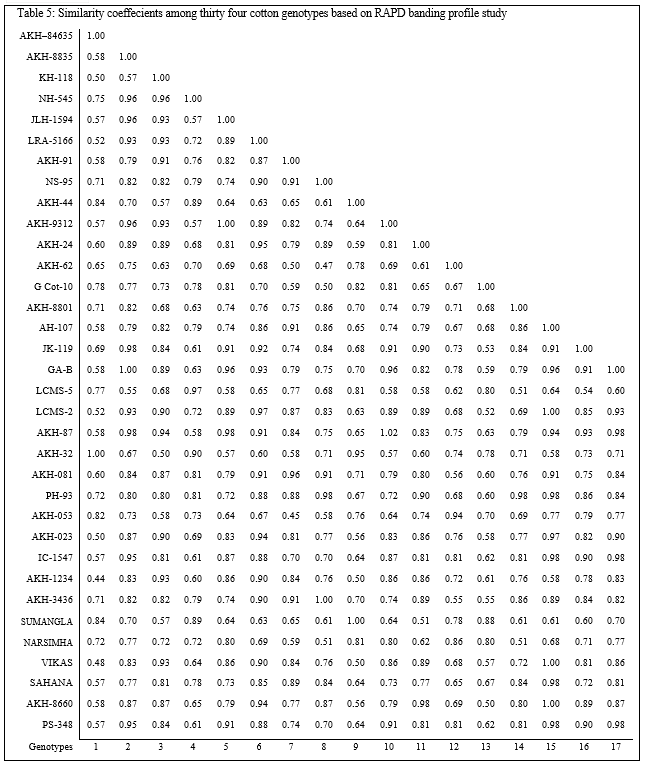
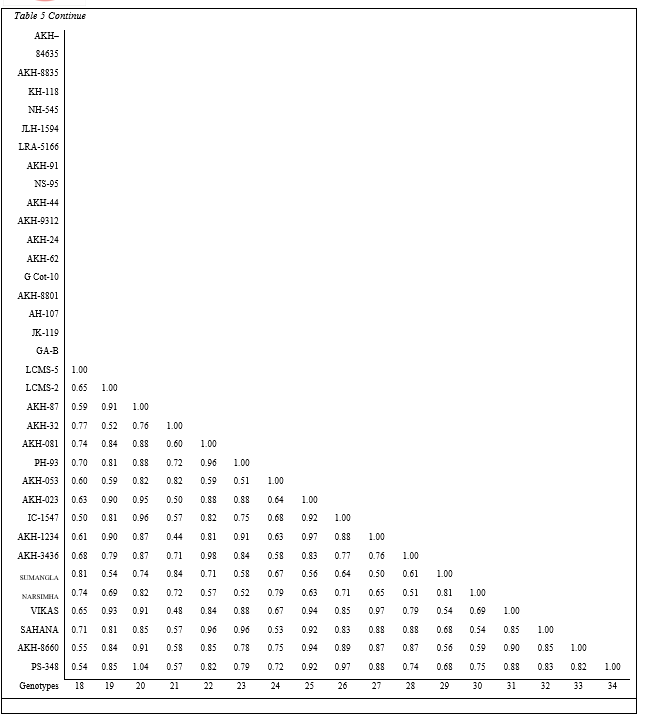
The cumulative data of similar and dissimilar bands obtained in five sets of primers were used for estimation of similarity matrix comprising indices for all possible combinations and presented inTable 5 (Plate 5). The similarity indices based on RAPD polymorphism among the genotypes of clusters ranged from 0.67 to 0.91. Also it can be concluded that the biochemical and molecular markers are adequate to judge the dissimilarity among the genotypes for quicker selection of parents with assumption that the banding profile is a measure of discrimination. The results for selection of parents based on biochemical and molecular marker techniques score high similarity with that of conventional D2 statistics technique.
The Protein, Peroxidase and Esterase studies showed more than fifty per cent similarity regarding selection of parents as proposed on the basis of morphological traits by employing Mahalanobis D2 Statistics. Whereas, RAPD and Polyphenol oxidase exhibited low similarity. Regarding the RAPD the results can be justified as the complete genome cannot be analyzed for polymorphism as the limited and random base segments were used. Hence for getting more precision in the predictions researchers have to use more number of random primers along with other advanced techniques to identify the genetic dissimilarity among the genotypes at molecular level.
Biochemical study further revealed that electrophoretic proteins and isozyme profiles could be efficiently used for distinguishing the genotypes. But the qualitative differences could not be reproduced as it is very difficult to identify and count the faint bands and needs expertise in discrimination based on presence or absence of bands. To overcome these limiting factors, the molecular marker analysis was undertaken. The Randomly Amplified Polymorphic DNA Markers (RAPDs) were used, which has also shown resemblance with the conventional method for categorization of genotypes.
IV. ACKNOWLEDGEMENT
Author is very much thankful to Senior Research Scientist, Cotton Research Unit and Head, Department of Agricultural Botany for providing seed material, field and laboratory facility during the present investigation.



Conclusion
The genetic diversity plays and important role in crop improvement program. The estimation of genetic distance gives an idea about the relationship of genotypes and their similarities and differences. Recently the biochemical and molecular marker techniques are found to be quicker for screening of large gene pool. Also it is helpful for selecting the genotypes for breeding program. These advanced biochemical and molecular techniques can now supplement to speed up the conventional breeding programs along with improvement in precision.
References
[1] AnuradhaVarier and MalvikaDadlani. 1992. Effect of ageing on profiles of soluble protein of cotton and esterase isozymes of Pearl millet seeds. Indian J. Plant Physiology.35 (2): 145 – 157. [2] Cherry, J.P.; F.R.H. Katterman and J.E.Endrizzi. 1970. Comparative studies of seed proteins of species of Gossypium by gel electrophoresis. Evolution,24 : 431 – 447. [3] Kharade. M.R., V.V.Ujjainkar and K.C.Bhgat. 2016. Evaluation of genetic diversity in groundnut mutatants (Arachishypogaea L.) using seed protein electrophoresis., Eco. Env. & Cons.22 : 83 – 87. [4] Krishna, T.G.andN.Jawali. 1997. DNA isolation from single or half seeds suitable for Random Amplified Polymorphic DNA analysis. Analytical Biochemistry, 250 : 125-127. [5] Mahalanobis, P.C. 1928. A statistical study of Chinese head measurement Man in India.,8 : 32 – 64. [6] Mahalanobis, P.C. 1936. On the generalized distance in statistics. Proc.National Institute. Sci. India 2 : 49 – 55. [7] MalvikaDadlani and Anuradha Varier. 1993. Electrophoresis for variety identification. Tech. Bulletin, IARI., New Delhi.:3 – 4. [8] Markert, C.L. and F. Moller. 1959. Multiple forms of enzymes: Tissue, ontogenetic and species - specific patterns. Proc. Natl. Acad. Sci., USA.,45 : 756 – 763. [9] Misagi, I.J. 1982. Physiology and Biochemistry of plant pathogen interactions. Plenurn Press., New York and London.: 110–163. [10] Nei., M. And W.H.Li. 1979. Mathematical model for studying genetic variation in terms of restriction indonucleases. Proc. Natl. Acad. Sci. USA. 76 (10):5269 – 5273. [11] Nerkar, Y.S. and T.N.Rao. 1993. Use of seed protein and enzymes polymorphism in the identification of cultivar of cotton.,Seed Res., 21 : 375 – 393. [12] Nijenhuis, B.T. 1971. Estimation of the proportion of inbred seed in Brussels sprouts hybrid seed by acid phosphate isozyme analysis. Euphytica.,20 : 498 – 507. [13] Ujjainkar V.V. and M.W.Marawar. 2020. Genetic Diversity: A Tool for Assessing Potential of Genepool., Remarking An Analisation.,4 (11): E38 - 44 [14] Ujjainkar V.V., S.B.Datke, D.G.Mahurkar and T.H.Rathod 2001. Dispersion study for annual volume production in teak (Tectona grandis) growing in natural and managed habitats., Annals of Plant Physiology 15 (1) : 80-84
Copyright
Copyright © 2022 Dr. Vaibhav V. Ujjainkar. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET45362
Publish Date : 2022-07-05
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

