Ijraset Journal For Research in Applied Science and Engineering Technology
- Home / Ijraset
- On This Page
- Abstract
- Introduction
- Conclusion
- References
- Copyright
Understanding Machine learning Applications in Cancer Prognosis and Early Detection
Authors: Kartikraj Shetty
DOI Link: https://doi.org/10.22214/ijraset.2022.40978
Certificate: View Certificate
Abstract
Machine learning and deep learning technologies have seen a recent growth in trend towards their applications in personalised and predictive medicine. The ML models are designed with an aim to supervise the progression of cancer within a patient and aid in its treatment. Cancer is quite a diverse condition, which means that an early diagnosis and timely screening is instrumental for treatment. An array of popular ML techniques is used in Cancer Prognosis including and not limited to Artificial Neural Networks, Decision Trees, Support Vector Machines, Bayesian Networks and other Deep Learning approaches. Each of these methodologies contribute to the development of predictive models. Each model developed is expected to substantially improve the accuracy of susception and recurrence prediction. However, a number of published studies that appear to have build over these models lack validation and/or appropriate testing. In this review, we analyse and present a view on recent development of ML approaches that are applied in Cancer Prognosis and Prediction modelling.
Introduction
I. INTRODUCTION
Currently the field of Artificial Intelligence and Machine Learning have gained traction and made remarkable advancements in multiple medical sectors. This has resulted in vast amounts of medical information and data being available for medical researchers. A number models based on ANN and DT’s have already been in use for over 20 years to aid with cancer diagnosis and type detection. SVM have recently made outstanding advancements and are considered to be the superior classifier of cancer prognosis when implemented in linear and non-linear problems. SVM based predictors are also proven to be better performers, however, their models are relatively immature.
ML models primarily are intended to aid in pattern identification and relation association on complex datasets. They also enable effective prediction outcomes of classified cancer type. In this paper we review several of these ML based predictive models and analyse the supervised techniques that they are based on. We will do so by analysing and studying the data that is integrated in each of the technologies and their respective performance in each proposition.
ML models incorporate an array of patient data that are retrieved from their clinical analyses such as family history, dietary habits, high-risk habits and environmental factors that the subject is exposed to like radiation through UV or other carcinogens. Each of these factors have played an evident role in predicting the subject’s risk for developing cancer (Leenhouts 1999; Cascon et al. 2004). Factors like the patients own genetic make-up and details that are obtained on a molecular level are also necessary inputs for any successful detection model. (Colozza et al. 2005)
As the number of parameters grow, the more challenging it becomes to develop a flexible and efficient detection or prognosis model. In this review, we study the challenges facing each of the models and try to establish the bases of their limitations. In order to do so, we must first understand the primary objective of any prognosis model. A prognosis model differs from a detection model as its fundamental goal is to predict the susceptibility, reoccurrence and survivability. The prognosis model relies heavily on the success and quality of the diagnosis. Hence medical diagnosis remains instrumental for any predictive prognosis to be carried out successfully.
II. MACHINE LEARNING METHODOLOGY
Machine learning, often confused with Artificial Intelligence is actually a subset or branch of AI that at its core uses statistically optimisation to learn from data samples. These learning are then used for probabilistic predictions for (i) calculating the unknown dependencies within any selected dataset of a sample system and (ii) predict their potential outcomes by using these calculated estimations. In simple terminology we can say that ML techniques are used to learn and classify data from complex datasets. The learning approach encompasses two methodologies namely supervised and unsupervised learning. Both of which have different procedural patterns and varied purposes respectively.
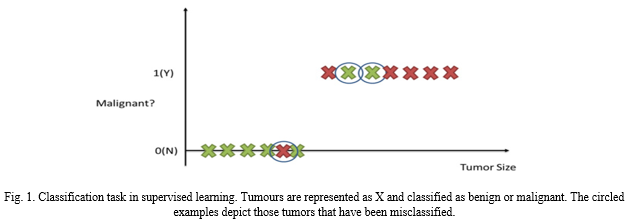
In supervised learning we device a dataset from which we draw desired outputs based on the input data that is fed. The learning model interprets and learns from the dataset, whereas in unsupervised learning, the dataset has no labelled data and the output completely depends on the model and its findings. There is no expectation of a certain output or any output at all. Clustering for example is a popular unsupervised learning task where we categories clusters based on their identified characteristics with no labelling of input data. The approach identifies and maps raw, unlabelled data into clusters of classifiable data. Therefore, allowing the learning model to find patterns and discover any potential grouping in the sample. Another form of ML method is semi-supervised learning where the dataset comprises of a combination of primarily unlabelled data with some labelled data present alongside it.
We estimate the predictive value of each new sample and the learning function maps these variables in real-value variables. However, within datasets with non-linear variables when the relationships are interdependent or partially dependent, statistical analysis tend to fall flat. Since any biological system is most often non-linear, a statistical approach is impossible. This is where ML algorithms tend to outshine statistical analysis.
ML however, is not perfect. Nor is it accurate every single time. Ever ML model has its drawback and limitations. These issues are caused by the quality of data provided. For example, a sample dataset may contain noise, outliers or redundancy issues that may lead to inconclusive results. As the size of dataset increases, these issues also scale and lead to challenges in designing an efficient model. This means that the quality of the dataset needs to be improved before one can implement any learning-based algorithms on the sample set. Processing of the dataset is essential so that the modified data may be better analysed. One such processing could be dimensionality reduction which can reduced the number of redundancy and irrelevant data enabling us to develop more robust learning models.
The primary goal of any ML techniques is to design a model that can not only provide classification of the dataset, but also enable us to perform prediction and estimation problems on the output set variables.
III. ML METHODS USED IN CANCER RESEARCH
Post processing of the data, we evaluate the data and determine the method that best defines our learning task. Common ML techniques used for Cancer Prognosis are (i) Artificial Neural Networks (ANN), (ii) Decision Trees (DT’s), (iii) Bayesian Networks (BN) and (iv) Support Vector Machines (SVM).
A. Artificial Neural Network
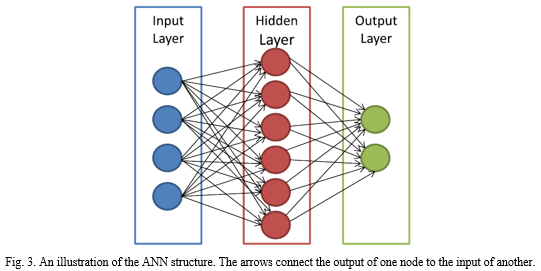
An ANN is trained to generate multiple combinations of outputs from the input variables which allows it to handle multiple problems pertaining to pattern recognition. Originally, the first Neural Network was designed to recreate the functioning of neurons in our brains and study how the interconnection between these neurons was carried though the axon junction. Neural Networks are designed on a layer analogy. A layer is a weight matrix that represents the wiring of interconnected neurons. Each layer would theoretically process an input and generate appropriate output in a string or vector based mathematical structure. The key challenge when using ANN for cancer detection is mapping real world input values into computer understandable numeric value or vector. For example, it is difficult to map a physical characteristic of a subject or their gene name type into a numeric vector.
Notably, the layered structure has been observed to take longer to process and yet have inadequate performance. The layered structure being unsupervised has also made inconclusive results hard to understand, as there is no way of knowing how the ANN arrived at this outcome. Thus, making it next to impossible to decipher any failed output. In other words, ANN has no feedback and has to be optimized individually for every application.
B. Support Vector Machine
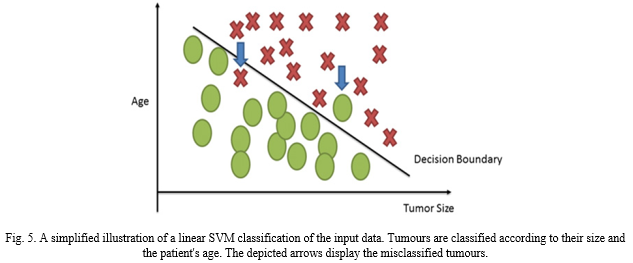
An SVM is designed for performing non-linear classification and it does so by creating a hyperplane. This hyperplane essential functions as a separator between two maximum margin classes. As a result, the distance between both the hyperplane itself and the margin is effectively maximised. SVM machines use what we call non-linear kernel. These kernels significantly improve the performance of the classification model built by the SVM. Similar to ANN, Support Vectors have great potential in the field of complex analysis. Be it handwriting recognition or text-based recognition. In medical diagnosis, it can help identify the protein function. SVMs can aid in the classification of tumours as they provide probabilistic output that can identify among malignant and benign tumour. SVMs also have a decision boundary that enables researchers to detect any misclassification that may have resulted by the method.
C. Decision Tree
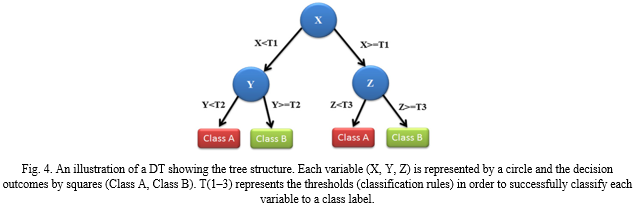
Decision tree as the name suggests represent a tree like structure that encompasses input variables represented as nodes and their respective variable outcomes labelled and represented as leaves. The Tree branches out into a scheme that identifies and visually represents corresponding outcomes as per their classification. Decision trees have been around for centuries and are the most common classification ML methods. They’re easy and simple to interpret and their models are quick to learn. The representation through decision trees allows for easy traversal across its classified branches thus allowing for a sample to be easily associated with its respective class. The Decision Tree architecture allows for easy reasoning and interpretation.
IV. SURVEY OF ML APPLICATIONS IN CANCER
From our analysis of the literature several trends were noted. As has been remarked previously, the use of machine learning in cancer prediction and prognosis is growing rapidly, with the number of papers increasing by 25% per year. These queries yielded 1061 and 157 hits respectively, giving a non-overlapping set of 1174 papers. Removing the 53 papers with machine learning components in this set, we were left with 1121 papers. While a detailed review of each abstract was not possible, a random sampling indicated that ~80% of these papers were relevant (890 papers) in that they used statistical approaches to predict or prognosticate cancer outcomes. When looking at the types of predictions or prognoses being made, the vast majority (86%) are associated with predicting cancer mortality (44%) and cancer recurrence (42%). However, a growing number of more recent studies are now aimed at predicting the occurrence of cancer or the risk factors associated with developing cancer. As a general rule, regardless of the machine learning method used, the type of prediction being made or the type of cancer being evaluated, machine learning methods appear to improve the accuracy of predictions by average of 15-25% over alternative or conventional approaches. Almost 70% of all reported studies use neural networks as their primary (and sometimes only) predictor. Support vector machines are a distant second with 9%, while clustering and decision trees each account for about 6%. However, a disturbing number of studies lacked sufficient internal or external validation, were trained on far too few examples, tested on only a single machine learner or had no well-defined standard with which to compare the performance of the reported algorithm.
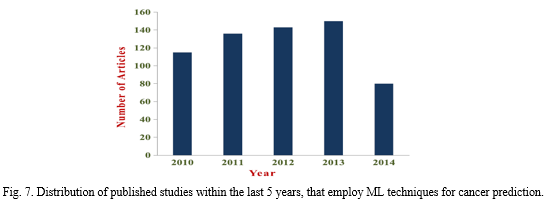
V. CASE STUDY ON PREDICTION OF CANCER RECURRENCE
The study of De Laurentiis et al. (1999), addresses some of the drawbacks noted in the previous studies. These authors aimed to predict the probability of relapse over a 5 years period for breast cancer patients. A combination of 7 prognostic variables was used including clinical data such as patient age, tumour size, and number of axillary metastases. Protein biomarker information such as oestrogen and progesterone receptor levels were also included.
The aim of the study was to develop an automatic, quantitative prognostic method that was more reliable than the classical tumour-node-metastasis (TNM) staging system. TNM is a physician-based expert system that relies heavily on the subjective opinion of a pathologist or expert clinician.
The authors employed an ANN-based model that used data from 2441 breast cancer patients (times 7 data points each) yielding a data set with more than 17,000 data points. This allowed the authors to maintain a sample-to-feature ratio of well over the suggested minimum of 5 (Somorjai et al. 2003).
The entire data set was partitioned into three equal groups: training (1/3), monitoring (1/3), and test sets (1/3) for optimization and validation. In addition, the authors also obtained a separate set of 310 breast cancer patient samples from a different institution, for external validation. This allowed the authors to assess the generalizability of their model outside their institution — a process not done by the two previously discussed studies.
This study is particularly notable not only for the quantity of data and the thoroughness of validation, but also for the level of quality assurance applied to the data handling and processing. For instance, the data was separately entered and stored in a relational database and all of it was independently verified by the referring physicians to maintain quality. With 2441 patients and 17,000 data points in the data set, the sample size was sufficiently large that a normal population distribution of breast cancer patients could be assumed within the data set, even after partitioning. Regardless, the authors explicitly verified this assumption by looking at the distribution of the data for the patients within each set (training, monitoring, test, and external) and showed that the distributions were relatively similar. This quality assurance and attention to detail allowed the authors to develop a very accurate and robust
classifier. Since the aim of the study was to develop a model that predicted relapse of breast cancer better than the classical TNM staging system, it was important for the ANN model to be compared to TNM staging predictions. This was done by comparing the performance using a receiver operator characteristic (ROC) curve. The ANN model (0.726) was found to outperform the TNM system (0.677) as measured by the area under the ROC curve.
This study is an excellent example of a well-designed and well tested application of machine learning. A sufficiently large data set was obtained and data for each sample was independently verified for quality assurance and accuracy. Furthermore, blinded sets for validation were available from both the original data set and from an external source to assess the generality of the machine learning model. Finally, the accuracy of the model was explicitly compared to that of a classical prognostic scheme, TNM staging. Perhaps the one drawback to this study was the fact that the authors only tested a single kind of machine learning (ANN) algorithm. Given the type and quantity of data used, it is quite possible that their ANN model may have been outperformed by another machine learning technique.
VI. DISCUSSION
Except the data size, the dataset quality as well as the careful feature selection schemes are of great importance for effective ML and subsequently for accurate cancer predictions. Choosing the most informative feature subset for training a model, by means of feature selection methods, could result in robust models. Additionally, feature sets that consist of histological or pathological assessments are characterized by reproducible values. Due to the lack of static entities when dealing with clinical variables it is important for a ML technique to be adjusted to different feature sets over time.
A key point to several studies, regarding their promising results, was the fact that several ML techniques were employed as an aim to find the most optimal one. Apart from this, the combination of multiple data types that would be fed as input to the models is also a trend. Looking back to the previous decade, only molecular and clinical information was exploited for making predictions of cancer outcomes. With the rapid development of HTTs, including genomic, proteomic and imaging technologies, new types of input parameters have been collected. We found that almost all the predictions were made by integrating either genomic, clinical, histological, imaging, demographic, epidemiological data and proteomic data or different combinations of these types.
A small sized training sample, compared to data dimensionality, can result in misclassifications while the estimators may produce unstable and biased models. It is obvious that a richer set of patients used for their survival prediction can enhance the generalizability of the predictive model.
Conclusion
In this review, we discussed the concepts of ML while we outlined their application in cancer prediction/prognosis. Most of the studies that have been proposed the last years and focus on the development of predictive models using supervised ML methods and classification algorithms aiming to predict valid disease outcomes. Based on the analysis of their results, it is evident that the integration of multidimensional heterogeneous data, combined with the application of different techniques for feature selection and classification can provide promising tools for inference in the cancer domain.
References
[1] Chao Tan, Hui Chen, Chengyun Xia, Early prediction of lung cancer based on the combination of trace element analysis in urine and an Adaboost algorithm, J. Pharm. Biomed. Anal. 49 (3) (2009) 746–752. [2] D.-H. Tae-WooKim, Chung-Yill Park, Decision tree of occupational lung cancer using classification and regression analysis, Safety Health Work 1 (2) (2010) 140–148. [3] M. Zie?ba, J.M. Tomczak, Marek Lubicz, Jerzy S´wia? tek, Boosted SVM for extracting rules from imbalanced data in application to prediction of the post-operative life expectancy in the lung cancer patients, Appl. Soft Comput. 14 (2014) 99–108. [4] Worrawat Engchuan, Jonathan H. Chan, Pathway activity transformation for multi-class classification of lung cancer datasets, Neurocomputing 165 (2015) 81–89. [5] H. Azzawi, J. Hou, Y. Xiang, R. Alanni, Lung cancer prediction from microarray data by gene expression programming, IET Syst. Biol. 10 (5) (2016) 168–178. [6] P. Petousis, S.X. Han, Denise Aberle, Alex A.T. Bui, Prediction of lung cancer incidence on the low-dose computed tomography arm of the National Lung Screening Trial: a dynamic Bayesian network, Artif. Intell. Med. 72 (2016) 42– 55. [7] C.M. Lynch, J.D. Behnaz Abdollahi, A. Fuqua, R. de Carlo, James A. Bartholomai, Rayeanne N. Balgemann, Victor H. van Berkel, Hermann B. Frieboes, Prediction of lung cancer patient survival via supervised machine learning classification techniques, Int. J. Med. Inf. 108 (2017) 1–8. [8] D.S. Rao, D.P. Tripathy, Optimization of machinery noise using Genetic Algorithm. Noise Conference 2017. Michigan, 2017; 527–537. [9] P. Petousis, A. Winter, W. Speier, D.R. Aberle, W. Hsu, A.A.T. Bui, Using sequential decision making to improve lung cancer screening performance, IEEE Access 7 (2019) 119403–119419. [10] V. Krishnaiah, G. Narsimha, C. Subhash, Diagnosis of lung cancer prediction system using data mining classification techniques, Int. J. Comp. Sci. Inf. Technol. 4 (1) (2013) 39–45. [11] T. Ojala, M. Pietikainen, T. Maenpaa, Multiresolution gray-scale and rotation invariant texture classification with local binary patterns, IEEE Trans. Pattern Anal. Mach. Intell. 24 (7) (2002) 971–987. [12] L. Demidova, I. Klyueva, Y. Sokolova, N. Stepanov, N. Tyart, Intellectual approaches to improvement of the classification decisions quality on the base of the SVM classifier, Procedia Comput. Sci. 103 (2017) 222–230. [13] N. Picco, R.A. Gatenby, A.R.A. Anderson, Stem cell plasticity and niche dynamics in cancer progression, IEEE Trans. Biomed. Eng. 64 (3) (2017) 528– 537. [14] Pawe? Krawczyk, Tomasz Kucharczyk, Kamila Wojas-Krawczyk, Screening of Gene Mutations in Lung Cancer for Qualification to Molecularly Targeted Therapies, INTECH Open Access Publisher, 2012. [15] A. Colquhoun, L. McHugh, E. Tulchinsky, M. Kriajevska, J. Mellon, Combination treatment with ionising radiation and Gefitinib (‘Iressa’, ZD1839), an epidermal growth factor receptor (EGFR) inhibitor, significantly inhibits bladder cancer cell growth in vitro and in vivo, J. Radiat. Res. 48 (5) (2007) 351–360. [16] E. Adetiba, O.O. Olugbara, Lung cancer prediction using neural network ensemble with histogram of oriented gradient genomic features, Sci. World J. (2015). [17] S.S. Alahmari, D. Cherezov, D.B. Goldgof, L.O. Hall, R.J. Gillies, M.B. Schabath, Delta radiomics improves pulmonary nodule malignancy prediction in lung cancer screening, IEEE Access 6 (2018) 77796–77806. [18] S. Park, S.J. Lee, E. Weiss, Y. Motai, Intra- and inter-fractional variation prediction of lung tumors using fuzzy deep learning, IEEE J. Transl. Eng. Health Med. 4 (2016) 1–12. [19] A. Raweh, M. Nassef, A. Badr, A hybridized feature selection and extraction approach for enhancing cancer prediction based on DNA methylation, IEEE Access 6 (2018) 15212–15223. [20] Delen D, Walker G, Kadam A. Predicting breast cancer survivability: a comparison of three data mining methods. Artif Intell Med 2005;34:113–27. [21] Urbanowicz RJ, Andrew AS, Karagas MR, Moore JH. Role of genetic heterogeneity and epistasis in bladder cancer susceptibility and outcome: a learning classifier sys tem approach. J Am Med Inform Assoc 2013;20:603–12. [22] Bochare A, Gangopadhyay A, Yesha Y, Joshi A, Yesha Y, Brady M, et al. Integrating do main knowledge in supervised machine learning to assess the risk of breast cancer. Int J Med Eng Inform 2014;6:87–99. [23] Gilmore S, Hofmann?Wellenhof R, Soyer HP. A support vector machine for decision support in melanoma recognition. Exp Dermatol 2010;19:830–5. [24] Mac Parthaláin N, Zwiggelaar R. Machine learning techniques and mammographic risk assessment. Digital mammography. Springer; 2010. pp. 664–672. [59] Hall MA. Feature selection for discrete and numeric class machine learning; 1999. [60] Kohavi R, John GH. Wrappers for feature subset selection. Artif Intell 1997;97: 273–324. [25] Estévez PA, Tesmer M, Perez CA, Zurada JM. Normalized mutual information feature selection. IEEE Trans Neural Netw 2009;20:189–201. [26] Barrett T, Troup DB, Wilhite SE, Ledoux P, Rudnev D, Evangelista C, et al. NCBI GEO: mining tens of millions of expression profiles — database and tools update. Nucleic Acids Res 2007;35:D760–5. [27] Niu Y, Otasek D, Jurisica I. Evaluation of linguistic features useful in extraction of interactions from PubMed; application to annotating known, high-throughput and predicted interactions in I2D. Bioinformatics 2010;26:111–9. [28] Howlader N, Noone A, Krapcho M, Garshell J, Neyman N, Aletkruse S. SEER Cancer Statistics Review, 1975–2010, [Online] National Cancer Institute. Bethesda, MD: Na tional Cancer Institute; 2013 [Online]. [29] Bian X, Klemm J, Basu A, Hadfield J, Srinivasa R, Parnell T, et al. Data submission and curation for caArray, a standard based microarray data repository system; 2009. [66] Papadopoulos A, Fotiadis DI, Costaridou L. Improvement of microcalcification cluster detection in mammography utilizing image enhancement techniques. Comput Biol Med 2008;38:1045–55. [30] Papadopoulos A, Fotiadis DI, Likas A. Characterization of clustered microcalcifications in digitized mammograms using neural networks and support vector machines. Artif Intell Med 2005;34:141–50. [31] Bilal E, Dutkowski J, Guinney J, Jang IS, Logsdon BA, Pandey G, et al. Improving breast cancer survival analysis through competition-based multidimensional modeling. PLoS Comput Biol 2013;9:e1003047. [32] Cuzick J, Dowsett M, Pineda S, Wale C, Salter J, Quinn E, et al. Prognostic value of a combined estrogen receptor, progesterone receptor, Ki-67, and human epidermal growth factor receptor 2 immunohistochemical score and comparison with the Ge nomic Health recurrence score in early breast cancer. J Clin Oncol 2011;29:4273–8. [33] Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol 2009;27:1160–7
Copyright
Copyright © 2022 Kartikraj Shetty. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.

Download Paper
Paper Id : IJRASET40978
Publish Date : 2022-03-25
ISSN : 2321-9653
Publisher Name : IJRASET
DOI Link : Click Here
 Submit Paper Online
Submit Paper Online

